Characterization of ssfR and ssgA, two genes involved in sporulation of Streptomyces griseus
- PMID: 10986257
- PMCID: PMC110997
- DOI: 10.1128/JB.182.19.5521-5529.2000
Characterization of ssfR and ssgA, two genes involved in sporulation of Streptomyces griseus
Abstract
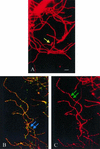
In the presence of cefoxitin, which inhibits septum formation during sporulation, Streptomyces griseus is unable to sporulate, retaining the sonication sensitivity of nonsporulating hyphae. Cefoxitin- and sonication-resistant mutant SKK2600 was isolated and showed many morphological differences from its parental strain. A 3.6-kb DNA fragment that complemented the mutations of SKK2600 contained two open reading frames (ORFs), either of which could complement SKK2600. One ORF, designated ssfR, encoded a protein containing a potential DNA-binding helix-turn-helix motif close to its N terminus. SsfR is similar to members of a large family of transcriptional regulators, particularly IclR of Escherichia coli. The second ORF was identified as ssgA, a previously described sporulation gene from S. griseus (S. Kawamoto and J. C. Ensign, Actinomycetology 9:136-151, 1995). A point mutation of C to T seven nucleotides upstream of the UGA stop codon of ssfR was responsible for the phenotype of isolated mutant strain SKK2600. Surprisingly, this mutation should not change the primary structure of SsfR. The ssfR and ssgA disruption mutants were constructed and showed the "white" mutant phenotype, with some growth medium dependence. In addition, the ssfR null mutant sporulated ectopically in phosphate starvation medium.
Figures

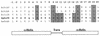



Similar articles
-
Transcriptional switch on of ssgA by A-factor, which is essential for spore septum formation in Streptomyces griseus.J Bacteriol. 2003 Feb;185(4):1273-83. doi: 10.1128/JB.185.4.1273-1283.2003. J Bacteriol. 2003. PMID: 12562798 Free PMC article.
-
Transcription of the sporulation gene ssgA is activated by the IclR-type regulator SsgR in a whi-independent manner in Streptomyces coelicolor A3(2).Mol Microbiol. 2004 Aug;53(3):985-1000. doi: 10.1111/j.1365-2958.2004.04186.x. Mol Microbiol. 2004. PMID: 15255907
-
A gene cluster involved in aerial mycelium formation in Streptomyces griseus encodes proteins similar to the response regulators of two-component regulatory systems and membrane translocators.J Bacteriol. 1993 Apr;175(7):2006-16. doi: 10.1128/jb.175.7.2006-2016.1993. J Bacteriol. 1993. PMID: 8458843 Free PMC article.
-
Expression analysis of the ssgA gene product, associated with sporulation and cell division in Streptomyces griseus.Microbiology (Reading). 1997 Apr;143 ( Pt 4):1077-1086. doi: 10.1099/00221287-143-4-1077. Microbiology (Reading). 1997. PMID: 9141673
-
Cloning and characterization of a gene involved in aerial mycelium formation in Streptomyces griseus.J Bacteriol. 1995 Nov;177(22):6401-10. doi: 10.1128/jb.177.22.6401-6410.1995. J Bacteriol. 1995. PMID: 7592414 Free PMC article.
Cited by
-
The Agrobacterium tumefaciens transcription factor BlcR is regulated via oligomerization.J Biol Chem. 2011 Jun 10;286(23):20431-40. doi: 10.1074/jbc.M110.196154. Epub 2011 Apr 4. J Biol Chem. 2011. PMID: 21467043 Free PMC article.
-
Expression of the melC operon in several Streptomyces strains is positively regulated by AdpA, an AraC family transcriptional regulator involved in morphological development in Streptomyces coelicolor.J Bacteriol. 2005 May;187(9):3180-7. doi: 10.1128/JB.187.9.3180-3187.2005. J Bacteriol. 2005. PMID: 15838045 Free PMC article.
-
Molecular control of gene expression by Brucella BaaR, an IclR-type transcriptional repressor.J Biol Chem. 2018 May 11;293(19):7437-7456. doi: 10.1074/jbc.RA118.002045. Epub 2018 Mar 22. J Biol Chem. 2018. PMID: 29567835 Free PMC article.
-
Constitutive expression of ftsZ overrides the whi developmental genes to initiate sporulation of Streptomyces coelicolor.Antonie Van Leeuwenhoek. 2012 Mar;101(3):619-32. doi: 10.1007/s10482-011-9678-7. Epub 2011 Nov 24. Antonie Van Leeuwenhoek. 2012. PMID: 22113698 Free PMC article.
-
Taxonomy, Physiology, and Natural Products of Actinobacteria.Microbiol Mol Biol Rev. 2015 Nov 25;80(1):1-43. doi: 10.1128/MMBR.00019-15. Print 2016 Mar. Microbiol Mol Biol Rev. 2015. PMID: 26609051 Free PMC article. Review.
References
-
- Champness W C, Chater K F. Regulation and integration of antibiotic production and morphological differentiation in Streptomyces spp. In: Piggot P Jr, Moran C P, Youngman P, editors. Regulation of bacterial differentiation. Washington, D.C.: American Society for Microbiology; 1994. pp. 61–94.
-
- Chater K F. Taking a genetic scalpel to the Streptomyces colony. Microbiology. 1998;144:1456–1478. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

