Cloning and nucleotide sequence of the DNA gyrase (gyrA) gene from Mycoplasma hominis and characterization of quinolone-resistant mutants selected in vitro with trovafloxacin
- PMID: 10991851
- PMCID: PMC90142
- DOI: 10.1128/AAC.44.10.2719-2727.2000
Cloning and nucleotide sequence of the DNA gyrase (gyrA) gene from Mycoplasma hominis and characterization of quinolone-resistant mutants selected in vitro with trovafloxacin
Abstract
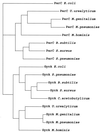
We report the cloning and characterization of the gyrA gene of the Mycoplasma hominis DNA gyrase, which was previously shown to be associated with quinolone resistance in this organism. The 2,733-bp gyrA gene encodes a protein of 911 amino acids with a calculated molecular mass of 102.5 kDa. As expected, M. hominis GyrA exhibits higher homology with the GyrA subunits of the gram-positive bacteria Clostridium acetobutylicum, Bacillus subtilis, Streptococcus pneumoniae, and Staphylococcus aureus than with its Escherichia coli counterpart. Knowing the entire sequence of the gyrA gene of M. hominis could be very useful for confirming the role of the GyrA subunit in fluoroquinolone resistance. Twenty-nine mutants of M. hominis were selected stepwise for resistance to trovafloxacin, a new potent fluoroquinolone, and their gyrA, gyrB, parC, and parE quinolone resistance-determining regions were characterized. Three rounds of selection yielded 3 first-step, 12 second-step, and 14 third-step mutants. The first-step mutants harbored a single substitution, Glu460-->Lys (E. coli coordinates), in ParE. GyrA changes, Ser83-->Leu, Glu87-->Lys, and Ala119-->Glu or Val, were found only in the second round of selection. At the third step, additional substitutions, at ParC Ser80, Ser81, and Glu84 and ParE Leu440, associated with high-level resistance to fluoroquinolones, appeared. Thus, high-level resistance to trovafloxacin required three steps and was associated with alterations in both fluoroquinolone targets. According to these genetic data, in M. hominis, as in Staphylococcus aureus and Streptococcus pneumoniae, topoisomerase IV seems to be the primary target of trovafloxacin.
Figures




Similar articles
-
Alterations in topoisomerase IV and DNA gyrase in quinolone-resistant mutants of Mycoplasma hominis obtained in vitro.Antimicrob Agents Chemother. 1998 Sep;42(9):2304-11. doi: 10.1128/AAC.42.9.2304. Antimicrob Agents Chemother. 1998. PMID: 9736554 Free PMC article.
-
Mutations in the gyrA, parC, and parE genes associated with fluoroquinolone resistance in clinical isolates of Mycoplasma hominis.Antimicrob Agents Chemother. 1999 Apr;43(4):954-6. doi: 10.1128/AAC.43.4.954. Antimicrob Agents Chemother. 1999. PMID: 10103208 Free PMC article.
-
[Role of mutations in parC and gyrA in forming resistance of Mycoplasma hominis to fluoroquinolones].Mol Gen Mikrobiol Virusol. 1999;(4):19-24. Mol Gen Mikrobiol Virusol. 1999. PMID: 10621934 Russian.
-
DNA gyrase and topoisomerase IV are dual targets of clinafloxacin action in Streptococcus pneumoniae.Antimicrob Agents Chemother. 1998 Nov;42(11):2810-6. doi: 10.1128/AAC.42.11.2810. Antimicrob Agents Chemother. 1998. PMID: 9797208 Free PMC article.
-
Navigating fluoroquinolone resistance in Gram-negative bacteria: a comprehensive evaluation.JAC Antimicrob Resist. 2024 Aug 14;6(4):dlae127. doi: 10.1093/jacamr/dlae127. eCollection 2024 Aug. JAC Antimicrob Resist. 2024. PMID: 39144447 Free PMC article. Review.
Cited by
-
Mutation rate and evolution of fluoroquinolone resistance in Escherichia coli isolates from patients with urinary tract infections.Antimicrob Agents Chemother. 2003 Oct;47(10):3222-32. doi: 10.1128/AAC.47.10.3222-3232.2003. Antimicrob Agents Chemother. 2003. PMID: 14506034 Free PMC article.
-
Molecular mechanism of fluoroquinolones resistance in Mycoplasma hominis clinical isolates.Braz J Microbiol. 2014 May 19;45(1):239-42. doi: 10.1590/s1517-83822014000100034. eCollection 2014. Braz J Microbiol. 2014. PMID: 24948939 Free PMC article.
-
Synthesis and biological evaluation of 3-(1,3,4-oxadiazol-2-yl)-1,8-naphthyridin-4(1H)-ones as cisplatin sensitizers.Medchemcomm. 2018 Sep 25;9(11):1949-1960. doi: 10.1039/c8md00464a. eCollection 2018 Nov 1. Medchemcomm. 2018. PMID: 30568762 Free PMC article.
-
The Expanding Role of Pyridine and Dihydropyridine Scaffolds in Drug Design.Drug Des Devel Ther. 2021 Oct 13;15:4289-4338. doi: 10.2147/DDDT.S329547. eCollection 2021. Drug Des Devel Ther. 2021. PMID: 34675489 Free PMC article. Review.
-
Contribution of topoisomerase IV mutation to quinolone resistance in Mycoplasma genitalium.Antimicrob Agents Chemother. 2013 Apr;57(4):1772-6. doi: 10.1128/AAC.01956-12. Epub 2013 Jan 28. Antimicrob Agents Chemother. 2013. PMID: 23357772 Free PMC article.
References
-
- Bébéar C, Robertson J. Determination of minimal inhibitory concentration. In: Tully J G, Razin S, editors. Molecular and diagnostic procedures in mycoplasmology. II. San Diego, Calif: Academic Press, Inc.; 1996. pp. 189–199.
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Research Materials

