Distinct properties of Mycobacterium tuberculosis single-stranded DNA binding protein and its functional characterization in Escherichia coli
- PMID: 11000276
- PMCID: PMC110771
- DOI: 10.1093/nar/28.19.3823
Distinct properties of Mycobacterium tuberculosis single-stranded DNA binding protein and its functional characterization in Escherichia coli
Abstract
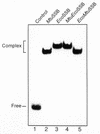
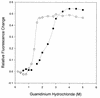
Single-stranded DNA binding proteins (SSBs) play an essential role in various DNA functions. Characterization of SSB from Mycobacterium tuberculosis, which infects nearly one-third of the world's population and kills about 2-3 million people every year, showed that its oligomeric state and various in vitro DNA binding properties were similar to those of the SSB from Escherichia coli. In this study, use of the yeast two-hybrid assay suggests that the ECO:SSB and the MTU:SSB are even capable of heterooligomerization. However, the MTU:SSB failed to complement a Deltassb strain of E. coli. The sequence comparison suggested that MTU:SSB contained a distinct C-terminal domain. The C-terminal domain of ECO:SSB interacts with various cellular proteins. The chimeric constructs between the N- and C-terminal domains of the MTU:SSB and ECO:SSB exist as homotetramers and demonstrate DNA binding properties similar to the wild-type counterparts. Despite similar biochemical properties, the chimeric SSBs also failed to complement the Deltassb strain of E.coli. These data allude to the occurrence of a 'cross talk' between the N- and the C-terminal domains of the SSBs for their in vivo function. Further, compared with those of the ECO:SSB, the secondary/tertiary interactions within MTU:SSB were found to be less susceptible to disruption by guanidinium hydrochloride. Such structural differences could be exploited for utilizing such essential proteins as crucial molecular targets for controlling the growth of the pathogen.
Figures





Similar articles
-
A genetic analysis of the functional interactions within Mycobacterium tuberculosis single-stranded DNA binding protein.PLoS One. 2014 Apr 10;9(4):e94669. doi: 10.1371/journal.pone.0094669. eCollection 2014. PLoS One. 2014. PMID: 24722671 Free PMC article.
-
Chimeras of Escherichia coli and Mycobacterium tuberculosis single-stranded DNA binding proteins: characterization and function in Escherichia coli.PLoS One. 2011;6(12):e27216. doi: 10.1371/journal.pone.0027216. Epub 2011 Dec 12. PLoS One. 2011. PMID: 22174737 Free PMC article.
-
Biochemical properties of single-stranded DNA-binding protein from Mycobacterium smegmatis, a fast-growing mycobacterium and its physical and functional interaction with uracil DNA glycosylases.J Mol Biol. 2002 May 17;318(5):1251-64. doi: 10.1016/s0022-2836(02)00053-0. J Mol Biol. 2002. PMID: 12083515
-
Escherichia coli single-stranded DNA-binding protein: multiple DNA-binding modes and cooperativities.Annu Rev Biochem. 1994;63:527-70. doi: 10.1146/annurev.bi.63.070194.002523. Annu Rev Biochem. 1994. PMID: 7979247 Review.
-
The mechanism of action of the SSB interactome reveals it is the first OB-fold family of genome guardians in prokaryotes.Protein Sci. 2021 Sep;30(9):1757-1775. doi: 10.1002/pro.4140. Epub 2021 Jun 14. Protein Sci. 2021. PMID: 34089559 Free PMC article. Review.
Cited by
-
A genetic analysis of the functional interactions within Mycobacterium tuberculosis single-stranded DNA binding protein.PLoS One. 2014 Apr 10;9(4):e94669. doi: 10.1371/journal.pone.0094669. eCollection 2014. PLoS One. 2014. PMID: 24722671 Free PMC article.
-
Chimeras of Escherichia coli and Mycobacterium tuberculosis single-stranded DNA binding proteins: characterization and function in Escherichia coli.PLoS One. 2011;6(12):e27216. doi: 10.1371/journal.pone.0027216. Epub 2011 Dec 12. PLoS One. 2011. PMID: 22174737 Free PMC article.
-
Analysis of the impact of a uracil DNA glycosylase attenuated in AP-DNA binding in maintenance of the genomic integrity in Escherichia coli.Nucleic Acids Res. 2010 Apr;38(7):2291-301. doi: 10.1093/nar/gkp1210. Epub 2010 Jan 7. Nucleic Acids Res. 2010. PMID: 20056657 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

