Messages through bottlenecks: on the combined use of slow and fast evolving polymorphic markers on the human Y chromosome
- PMID: 11023811
- PMCID: PMC1288547
- DOI: 10.1016/S0002-9297(07)62935-8
Messages through bottlenecks: on the combined use of slow and fast evolving polymorphic markers on the human Y chromosome
Figures


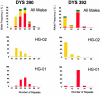
References
Electronic-Database Information
-
- BUGS Analyses, http://www.maths.abdn.ac.uk/~ijw/ (for BATWING)
References
-
- Bandelt H-J, Forster P, Röhl A (1999) Median-joining networks for inferring intraspecific phylogenies. Mol Biol Evol 16:37–48 - PubMed
-
- Bertranpetit J, Calafell F (1996) Genetic and geographic variation in cystic fibrosis: evolutionary considerations. In: Chadwick D, Cardew G (eds) Variation in the human genome. John Wiley & Sons, New York, pp 97–118
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

