Regulation of ferritin-mediated cytoplasmic iron storage by the ferric uptake regulator homolog (Fur) of Helicobacter pylori
- PMID: 11029412
- PMCID: PMC94726
- DOI: 10.1128/JB.182.21.5948-5953.2000
Regulation of ferritin-mediated cytoplasmic iron storage by the ferric uptake regulator homolog (Fur) of Helicobacter pylori
Abstract
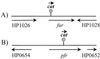
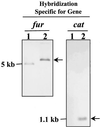
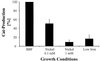
Homologs of the ferric uptake regulator Fur and the iron storage protein ferritin play a central role in maintaining iron homeostasis in bacteria. The gastric pathogen Helicobacter pylori contains an iron-induced prokaryotic ferritin (Pfr) which has been shown to be involved in protection against metal toxicity and a Fur homolog which has not been functionally characterized in H. pylori. Analysis of an isogenic fur-negative mutant revealed that H. pylori Fur is required for metal-dependent regulation of ferritin. Iron starvation, as well as medium supplementation with nickel, zinc, copper, and manganese at nontoxic concentrations, repressed synthesis of ferritin in the wild-type strain but not in the H. pylori fur mutant. Fur-mediated regulation of ferritin synthesis occurs at the mRNA level. With respect to the regulation of ferritin expression, Fur behaves like a global metal-dependent repressor which is activated under iron-restricted conditions but also responds to different metals. Downregulation of ferritin expression by Fur might secure the availability of free iron in the cytoplasm, especially if iron is scarce or titrated out by other metals.
Figures
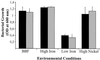



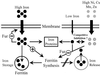
References
-
- Alm R A, Ling L S, Moir D T, King B L, Brown E D, Doig P C, Smith D R, Noonan B, Guild B C, deJonge B L, Carmel G, Tummino P J, Caruso A, Uria-Nickelsen M, Mills D M, Ives C, Gibson R, Merberg D, Mills S D, Jiang Q, Taylor D E, Vovis G F, Trust T J. Genomic-sequence comparison of two unrelated isolates of the human gastric pathogen Helicobacter pylori. Nature. 1999;397:176–180. - PubMed
-
- Althaus E W, Outten C E, Olson K E, Cao H, O'Halloran T V. The Ferric Uptake Regulation (Fur) repressor is a zinc metalloprotein. Biochemistry. 1999;38:6559–6569. - PubMed
-
- Andrews S C. Iron storage in bacteria. Adv Microb Physiol. 1998;40:281–351. - PubMed
-
- Ausubel F M, Brent R, Kingston R E, Moore D D, Seidman J G, Smith J A, Struhl K. Short protocols in molecular biology. New York, N.Y: John Wiley & Sons; 1992.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

