Life cycle of an endogenous retrovirus, ZAM, in Drosophila melanogaster
- PMID: 11044110
- PMCID: PMC110940
- DOI: 10.1128/jvi.74.22.10658-10669.2000
Life cycle of an endogenous retrovirus, ZAM, in Drosophila melanogaster
Abstract
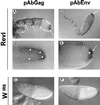
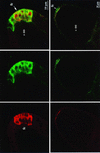
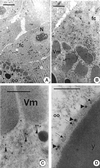
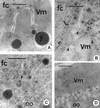
ZAM is an env-containing member of the gypsy family of retrotransposons that represents a possible retrovirus of invertebrates. In this paper, we traced ZAM mobilization to get information about a potential path a retroelement may take to reach the germ line of its host. In situ hybridization on whole-mount tissues and immunocytochemistry analyses with antibodies raised against ZAM Gag and Env proteins have shown that all components necessary to assemble ZAM viral particles, i.e., ZAM full-length RNAs and Gag and Env polypeptides, are coexpressed in a small set of follicle cells surrounding the oocyte. By electron microscopy, we have shown that ZAM viral particles are indeed detected in this somatic lineage of cells, which they leave and enter the closely apposed oocyte. Our data provide evidence that the vesicular traffic and yolk granules in the process of vitellogenesis play an important role in ZAM transfer to the oocyte. Our data support the possibility that vitellogenin transfer to the oocyte may help a retroelement pass to the germ line with no need of its envelope product.
Figures




References
-
- Coffin J M, Hughes S H, Varmus H E. Retroviruses. Cold Spring Harbor, N.Y: Cold Spring Harbor Laboratory Press; 1997. - PubMed
-
- Desset S, Conte C, Dimitri P, Calco V, Dastugue B, Vaury C. Mobilization of two retroelements, ZAM and Idefix, in a novel unstable line of Drosophila melanogaster. Mol Biol Evol. 1999;16:54–66. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

