Spore photoproduct (SP) lyase from Bacillus subtilis specifically binds to and cleaves SP (5-thyminyl-5,6-dihydrothymine) but not cyclobutane pyrimidine dimers in UV-irradiated DNA
- PMID: 11053385
- PMCID: PMC94787
- DOI: 10.1128/JB.182.22.6412-6417.2000
Spore photoproduct (SP) lyase from Bacillus subtilis specifically binds to and cleaves SP (5-thyminyl-5,6-dihydrothymine) but not cyclobutane pyrimidine dimers in UV-irradiated DNA
Abstract
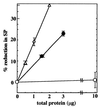
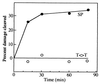
The predominant photolesion in the DNA of UV-irradiated dormant bacterial spores is the thymine dimer 5-thyminyl-5,6-dihydrothymine, commonly referred to as spore photoproduct (SP). A major determinant of SP repair during spore germination is its direct reversal by the enzyme SP lyase, encoded by the splB gene in Bacillus subtilis. SplB protein containing an N-terminal tag of six histidine residues [(6His)SplB] was purified from dormant B. subtilis spores and shown to efficiently cleave SP but not cyclobutane cis,syn thymine-thymine dimers in vitro. In contrast, SplB protein containing an N-terminal 10-histidine tag [(10His)SplB] purified from an Escherichia coli overexpression system was incompetent to cleave SP unless the 10-His tag was first removed by proteolysis at an engineered factor Xa site. To assay the parameters of binding of SplB protein to UV-damaged DNA, a 35-bp double-stranded oligonucleotide was constructed which carried a single pair of adjacent thymines on one strand. Irradiation of the oligonucleotide in aqueous solution or at 10% relative humidity resulted in formation of cyclobutane pyrimidine dimers (Py lozengePy) or SP, respectively. (10His)SplB was assayed for oligonucleotide binding using a DNase I protection assay. In the presence of (10His)SplB, the SP-containing oligonucleotide was selectively protected from DNase I digestion (half-life, >60 min), while the Py lozengePy-containing oligonucleotide and the unirradiated oligonucleotide were rapidly digested by DNase I (half-lives, 6 and 9 min, respectively). DNase I footprinting of (10His)SplB bound to the artificial substrate was carried out utilizing the (32)P end-labeled 35-bp oligonucleotide containing SP. DNase I footprinting showed that SplB protected at least a 9-bp region surrounding SP from digestion with DNase I with the exception of two DNase I-hypersensitive sites within the protected region. (10His)SplB also caused significant enhancement of DNase I digestion of the SP-containing oligonucleotide for at least a full helical turn 3' to the protected region. The data suggest that binding of SP lyase to SP causes significant bending or distortion of the DNA helix in the vicinity of the lesion.
Figures


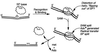
References
-
- Donnellan J E, Jr, Setlow R B. Thymine photoproducts but not thymine dimers are found in ultraviolet irradiated bacterial spores. Science. 1965;149:308–310. - PubMed
-
- Friedberg E C, Walker G C, Siede W. DNA repair and mutagenesis. Washington, D.C.: ASM Press; 1995.
-
- Gross D S, English K E, Collins K W, Lee S. Genomic footprinting of the yeast HSP82 promoter reveals marked distortion of the DNA helix and constitutive occupancy of heat shock and TATA elements. J Mol Biol. 1990;216:611–632. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

