Identification of an ancillary protein, YabF, required for activity of the KefC glutathione-gated potassium efflux system in Escherichia coli
- PMID: 11053405
- PMCID: PMC94807
- DOI: 10.1128/JB.182.22.6536-6540.2000
Identification of an ancillary protein, YabF, required for activity of the KefC glutathione-gated potassium efflux system in Escherichia coli
Abstract
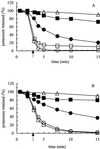
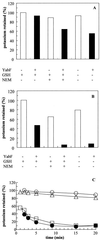
A new subunit, YabF, for the KefC K(+) efflux system in Escherichia coli has been identified. The subunit is required for maximum activity of KefC. Deletion of yabF reduces KefC activity 10-fold, and supply of YabF in trans restores activity. IS2 and IS10R insertions in yabF can be isolated as suppressors of KefC activity consequent upon the V427A and D264A KefC mutations.
Figures

Similar articles
-
Mutations in the glutathione-gated KefC K+ efflux system of Escherichia coli that cause constitutive activation.J Biol Chem. 1997 Oct 3;272(40):24942-7. doi: 10.1074/jbc.272.40.24942. J Biol Chem. 1997. PMID: 9312097
-
The K(+)-efflux system, KefC, in Escherichia coli: genetic evidence for oligomeric structure.Mol Membr Biol. 1994 Jan-Mar;11(1):55-61. doi: 10.3109/09687689409161030. Mol Membr Biol. 1994. PMID: 8019602
-
Different foci for the regulation of the activity of the KefB and KefC glutathione-gated K+ efflux systems.J Biol Chem. 1999 Apr 2;274(14):9524-30. doi: 10.1074/jbc.274.14.9524. J Biol Chem. 1999. PMID: 10092637
-
Methylglyoxal production in bacteria: suicide or survival?Arch Microbiol. 1998 Oct;170(4):209-18. doi: 10.1007/s002030050635. Arch Microbiol. 1998. PMID: 9732434 Review.
-
Protective mechanisms against toxic electrophiles in Escherischia coli.Trends Microbiol. 1999 Jun;7(6):242-7. doi: 10.1016/s0966-842x(99)01510-3. Trends Microbiol. 1999. PMID: 10366861 Review.
Cited by
-
Conserved and diversified gene families of monovalent cation/h(+) antiporters from algae to flowering plants.Front Plant Sci. 2012 Feb 14;3:25. doi: 10.3389/fpls.2012.00025. eCollection 2012. Front Plant Sci. 2012. PMID: 22639643 Free PMC article.
-
KefF, the regulatory subunit of the potassium efflux system KefC, shows quinone oxidoreductase activity.J Bacteriol. 2011 Sep;193(18):4925-32. doi: 10.1128/JB.05272-11. Epub 2011 Jul 8. J Bacteriol. 2011. PMID: 21742892 Free PMC article.
-
The Potassium Efflux System Kef: Bacterial Protection against Toxic Electrophilic Compounds.Membranes (Basel). 2023 Apr 27;13(5):465. doi: 10.3390/membranes13050465. Membranes (Basel). 2023. PMID: 37233526 Free PMC article. Review.
-
Regulation of ion channels by pyridine nucleotides.Circ Res. 2013 Feb 15;112(4):721-41. doi: 10.1161/CIRCRESAHA.111.247940. Circ Res. 2013. PMID: 23410881 Free PMC article. Review.
-
Untargeted metabolomics links glutathione to bacterial cell cycle progression.Nat Metab. 2020 Feb;2(2):153-166. doi: 10.1038/s42255-019-0166-0. Epub 2020 Feb 3. Nat Metab. 2020. PMID: 32090198 Free PMC article.
References
-
- Ausubel F M, Brent R, Kingston R E, Moore D D, Smith J A, Sideman J G, Struhl K. Current protocols in molecular biology. New York, N.Y: John Wiley and Sons, Inc.; 1987.
-
- Booth I R, Epstein W, Giffard P M, Rowland G C. Roles of the trkB and trkC gene products of Escherichia coli in K+ transport. Biochimie. 1985;67:83–90. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

