Succession of pelagic marine bacteria during enrichment: a close look at cultivation-induced shifts
- PMID: 11055904
- PMCID: PMC92360
- DOI: 10.1128/AEM.66.11.4634-4640.2000
Succession of pelagic marine bacteria during enrichment: a close look at cultivation-induced shifts
Abstract
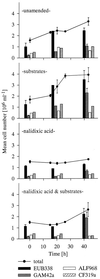
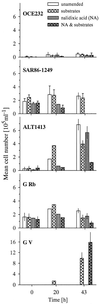
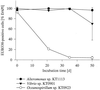
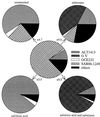
Enrichment experiments with North Sea bacterioplankton were performed to test if rapid incubation-induced changes in community structure explain the frequent isolation of members of a few particular bacterial lineages or if readily culturable bacteria are common in the plankton but in a state of dormancy. A metabolic inhibitor of cell division (nalidixic acid [NA]) was added to substrate-amended (S+) and unamended (S-) grazer-free seawater samples, and shifts in community composition and per cell DNA and protein content were compared with untreated controls. In addition, starvation survival experiments were performed on selected isolates. Incubations resulted in rapid community shifts towards typical culturable genera rather than in the activation of either dormant cells or the original DNA-rich bacterial fraction. Vibrio spp. and members of the Alteromonas/Colwellia cluster (A/C) were selectively enriched in S+ and S-, respectively, and this trend was even magnified by the addition of NA. These increases corresponded with the rise of cell populations with distinctively different but generally higher protein and DNA content in the various treatments. Uncultured dominant gamma-proteobacteria affiliating with the SAR86 cluster and members of the culturable genus Oceanospirillum were not enriched or activated, but there was no indication of substrate-induced cell death, either. Strains of Vibrio and A/C maintained high ribosome levels in pure cultures during extended periods of starvation, whereas Oceanospirillum spp. did not. The life strategy of rapidly enriched culturable gamma-proteobacteria could thus be described as a "feast and famine" existence involving different activation levels of substrate concentration.
Figures

References
-
- Albertini S, Chetelat A A, Miller B, Muster W, Pujadas E, Strobel R, Gocke E. Genotoxicity of 17 gyrase and four mammalian topoisomerase II poisons in prokaryotic and eukaryotic test systems. Mutagenesis. 1995;10:343–351. - PubMed
-
- Ammerman J W, Fuhrman J A, Hagstöm A, Azam F. Bacterioplankton growth in seawater. I. Growth kinetics and cellular characteristics in seawater cultures. Mar Ecol Prog Ser. 1984;18:31–39.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

