HIV-1 LTR as a target for synthetic ribozyme-mediated inhibition of gene expression: site selection and inhibition in cell culture
- PMID: 11058100
- PMCID: PMC113160
- DOI: 10.1093/nar/28.21.4059
HIV-1 LTR as a target for synthetic ribozyme-mediated inhibition of gene expression: site selection and inhibition in cell culture
Abstract
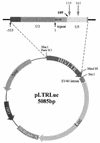
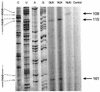
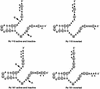
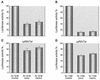
A library of three synthetic ribozymes with randomized arms, targeting NUX, GUX and NXG triplets, respectively, were used to identify ribozyme-accessible sites on the HIV-1 LTR transcript comprising positions -533 to 386. Three cleavable sites were identified at positions 109, 115 and 161. Ribozymes were designed against these sites, either unmodified or with 2'-modifications and phosphorothioate groups, and their cleavage activities of the transcript were determined. Their biological activities were assessed in cell culture, using a HIV-1 model assay system where the LTR is a promoter for the expression of the reporter gene luciferase in a transient expression system. Intracellular efficiency of the ribozymes were determined by cotransfection of ribozyme and plasmid DNA, expressing the target RNA. Modified ribozymes, directed against positions 115 and 161, lowered the level of LTR mRNA in the cell resulting in inhibition of expression of the LTR-driven reporter gene luciferase of 87 and 61%, respectively. In the presence of Tat the inhibitions were 43 and 25%. The inactive variants of these ribozymes exhibited a similar inhibitory effect. RNase protection revealed a reduction of RNA which was somewhat stronger for the active than the inactive ribozymes, particularly for ribozyme 115. Unmodified ribozymes showed no inhibition in the cell. The third ribozyme, targeting a GUG-triplet at position 109, possessed only low cleavage activity in vitro and no inhibitory effect in cell culture.
Figures


Similar articles
-
Secondary structure prediction and in vitro accessibility of mRNA as tools in the selection of target sites for ribozymes.Nucleic Acids Res. 2000 Nov 1;28(21):4113-24. doi: 10.1093/nar/28.21.4113. Nucleic Acids Res. 2000. PMID: 11058107 Free PMC article.
-
Inhibition of luciferase expression by synthetic hammerhead ribozymes and their cellular uptake.Nucleic Acids Res. 1999 Aug 1;27(15):3159-67. doi: 10.1093/nar/27.15.3159. Nucleic Acids Res. 1999. PMID: 10454613 Free PMC article.
-
Cleavage of an inaccessible site by the maxizyme with two independent binding arms: an alternative approach to the recruitment of RNA helicases.J Biochem. 2002 Jul;132(1):149-55. doi: 10.1093/oxfordjournals.jbchem.a003193. J Biochem. 2002. PMID: 12097172
-
The hairpin ribozyme: discovery, two-dimensional model, and development for gene therapy.Prog Nucleic Acid Res Mol Biol. 1998;58:1-39. doi: 10.1016/s0079-6603(08)60032-x. Prog Nucleic Acid Res Mol Biol. 1998. PMID: 9308362 Review.
-
Mechanism of action of hammerhead ribozymes and their applications in vivo: rapid identification of functional genes in the post-genome era by novel hybrid ribozyme libraries.Biochem Soc Trans. 2002 Nov;30(Pt 6):1145-9. doi: 10.1042/bst0301145. Biochem Soc Trans. 2002. PMID: 12440992 Review.
Cited by
-
Creating a ribonuclease T-tat that preferentially recognizes and hydrolyzes HIV-1 TAR RNA in vitro and in vivo.Nucleic Acids Res. 2008 Feb;36(3):963-9. doi: 10.1093/nar/gkm1118. Epub 2007 Dec 17. Nucleic Acids Res. 2008. PMID: 18086702 Free PMC article.
-
The activity of siRNA in mammalian cells is related to structural target accessibility: a comparison with antisense oligonucleotides.Nucleic Acids Res. 2003 Aug 1;31(15):4417-24. doi: 10.1093/nar/gkg649. Nucleic Acids Res. 2003. PMID: 12888501 Free PMC article.
-
A Conserved Target Site in HIV-1 Gag RNA is Accessible to Inhibition by Both an HDV Ribozyme and a Short Hairpin RNA.Mol Ther Nucleic Acids. 2014 Jul 29;3(7):e178. doi: 10.1038/mtna.2014.31. Mol Ther Nucleic Acids. 2014. PMID: 25072692 Free PMC article.
-
Identifying ribozyme-accessible sites using NUH triplet-targeting gapmers.Nucleic Acids Res. 2001 May 1;29(9):1906-14. doi: 10.1093/nar/29.9.1906. Nucleic Acids Res. 2001. PMID: 11328874 Free PMC article.
References
-
- Birikh K.R., Heaton,P.A. and Eckstein,F. (1997) The structure, function and application of the hammerhead ribozyme. Eur. J. Biochem., 245, 1–16. - PubMed
-
- Bramlage B., Luzi,E. and Eckstein,F. (1998) Designing ribozymes for the inhibition of gene expression. Trends Biotechnol., 16, 434–438. - PubMed
-
- James H.A. and Gibson,I. (1998) The therapeutic potential of ribozymes. Blood, 91, 371–382. - PubMed
-
- Muotri A.R., da Veiga Pereira,L., dos Reis Vasques,L. and Menck,C.F. (1999) Ribozymes and the anti-gene therapy: how a catalytic RNA can be used to inhibit gene function. Gene, 37, 303–310. - PubMed
-
- Lee C.G.L., Jeang,K.-T., Martin,M.A., Pastan,I. and Gotteman,M.M. (1997) Efficient long-term coexpression of a hammerhead ribozyme targeted to the U5 region of HIV-1 LTR by linkage to the multidrug-resistance gene. Antisense Nucleic Acid Drug Devel., 7, 511–522. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

