PCR hot start using primers with the structure of molecular beacons (hairpin-like structure)
- PMID: 11058144
- PMCID: PMC113163
- DOI: 10.1093/nar/28.21.e94
PCR hot start using primers with the structure of molecular beacons (hairpin-like structure)
Abstract
A new technique of PCR hot start using oligonucleotide primers with a stem-loop structure is developed here. The molecular beacon oligonucleotide structure without any chromophore addition to the ends was used. The 3'-end sequence of the primers was complementary to the target and five or six nucleotides complementary to the 3'-end were added to the 5'-end. During preparation of the reaction mixture and initial heating, the oligonucleotide has a stem-loop structure and cannot serve as an effective primer for DNA polymerase. After heating to the annealing temperature it acquires a linear structure and primer extension can begin.
Figures


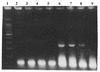
Similar articles
-
Controlled hot start and improved specificity in carrying out PCR utilizing touch-up and loop incorporated primers (TULIPS).Biotechniques. 2000 Nov;29(5):1018-20, 1022-4. doi: 10.2144/00295st03. Biotechniques. 2000. PMID: 11084864
-
Annealing control primer system for improving specificity of PCR amplification.Biotechniques. 2003 Dec;35(6):1180-4. doi: 10.2144/03356st03. Biotechniques. 2003. PMID: 14682052
-
Hot start PCR with heat-activatable primers: a novel approach for improved PCR performance.Nucleic Acids Res. 2008 Nov;36(20):e131. doi: 10.1093/nar/gkn575. Epub 2008 Sep 16. Nucleic Acids Res. 2008. PMID: 18796527 Free PMC article.
-
[Polymerase chain reaction, cold probes and clinical diagnosis].Sante. 1994 Jan-Feb;4(1):43-52. Sante. 1994. PMID: 7909267 Review. French.
-
Homogeneous detection of nucleic acids using self-quenched polymerase chain reaction primers labeled with a single fluorophore (LUX primers).Methods Mol Biol. 2006;335:95-114. doi: 10.1385/1-59745-069-3:95. Methods Mol Biol. 2006. PMID: 16785623 Review.
Cited by
-
Application of single-nucleotide polymorphism and mycobacterial interspersed repetitive units-variable number of tandem repeats analyses to clinical Mycobacterium tuberculosis isolates from Korea.Korean J Lab Med. 2011 Jan;31(1):37-43. doi: 10.3343/kjlm.2011.31.1.37. Korean J Lab Med. 2011. PMID: 21239869 Free PMC article.
-
Color-coded molecular beacons for multiplex PCR screening assays.PLoS One. 2019 Mar 18;14(3):e0213906. doi: 10.1371/journal.pone.0213906. eCollection 2019. PLoS One. 2019. PMID: 30883590 Free PMC article.
-
A new class of homogeneous nucleic acid probes based on specific displacement hybridization.Nucleic Acids Res. 2002 Jan 15;30(2):E5. doi: 10.1093/nar/30.2.e5. Nucleic Acids Res. 2002. PMID: 11788731 Free PMC article.
-
Novel hairpin-shaped primer assay to study the association of the -44 single-nucleotide polymorphism of the DEFB1 gene with early-onset periodontal disease.Clin Diagn Lab Immunol. 2004 Jul;11(4):766-9. doi: 10.1128/CDLI.11.4.766-769.2004. Clin Diagn Lab Immunol. 2004. PMID: 15242954 Free PMC article.
-
Homogeneous antibody-based proximity extension assays provide sensitive and specific detection of low-abundant proteins in human blood.Nucleic Acids Res. 2011 Aug;39(15):e102. doi: 10.1093/nar/gkr424. Epub 2011 Jun 6. Nucleic Acids Res. 2011. PMID: 21646338 Free PMC article.
References
-
- Horton R.M., Hoppe,B.L. and Conti-Tronconi,B.M. (1994) Biotechniques, 16, 42–43. - PubMed
-
- Bassam B.J. and Caetano-Anolles,G. (1993) Biotechniques, 14, 30–34. - PubMed
-
- Kellogg D.E., Rybalkin,I., Chen,S., Mukhamedova,N., Vlasik,T., Siebert,P.D. and Chenchik,A. (1994) Biotechniques, 16, 1134–1137. - PubMed
-
- Dang C. and Jayasena S.D., (1997) J. Mol. Biol., 264, 268–278. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

