Application of tRNA intergenic spacer PCR for identification of Enterococcus species
- PMID: 11060090
- PMCID: PMC87563
- DOI: 10.1128/JCM.38.11.4201-4207.2000
Application of tRNA intergenic spacer PCR for identification of Enterococcus species
Abstract
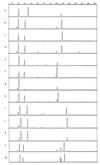
tRNA intergenic spacer PCR (tDNA-PCR) was evaluated for its usefulness in the differentiation of enterococcal species of human and animal origin. This technique was carried out for 124 strains belonging to 17 enterococcal species and generated DNA fragments, which were separated by capillary electrophoresis. tDNA-PCR enabled us to discriminate for all species tested. Enterococcus faecium showed minor but reproducible differences with Enterococcus durans, while Enterococcus hirae was easily distinguishable. Enterococcus avium, Enterococcus malodoratus, and Enterococcus raffinosus generated highly similar though distinctive patterns.
Figures

References
-
- Degheldre Y, Vandamme P, Goossens H, Struelens M. Identification of clinically relevant viridans streptococci by analysis of transfer DNA intergenic spacer length polymorphism. Int J Syst Bacteriol. 1999;49:1591–1598. - PubMed
-
- Descheemaeker P, Lammens C, Pot B, Vandamme P, Goossens H. Evaluation of arbitrarily primed PCR analysis and pulsed-field gel electrophoresis of large genomic DNA fragments for identification of enterococci important in human medicine. Int J Syst Bacteriol. 1997;47:555–561. - PubMed
-
- Devriese L A, Pot B, Collins M D. Phenotypic identification of the genus Enterococcus and differentiation of phylogenetically distinct enterococcal species and species groups. J Appl Bacteriol. 1993;75:399–408. - PubMed
-
- Devriese L A, Pot B, Kersters K, Lauwers S, Haesebrouck F. Acidification of methyl-α-d-glucopyranoside: a useful test to differentiate Enterococcus casseliflavus and Enterococcus gallinarum from Enterococcus faecium species group and from Enterococcus faecalis. J Clin Microbiol. 1996;34:2607–2608. - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

