The level of DAL80 expression down-regulates GATA factor-mediated transcription in Saccharomyces cerevisiae
- PMID: 11073899
- PMCID: PMC111397
- DOI: 10.1128/JB.182.23.6584-6591.2000
The level of DAL80 expression down-regulates GATA factor-mediated transcription in Saccharomyces cerevisiae
Abstract
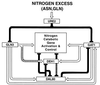
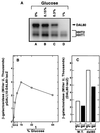
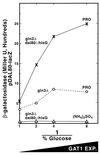
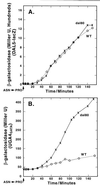
Nitrogen-catabolic gene expression in Saccharomyces cerevisiae is regulated by the action of four GATA family transcription factors: Gln3p and Gat1p/Nil1p are transcriptional activators, and Dal80 and Deh1p/Gzf3p are repressors. In addition to the GATA sequences situated upstream of all nitrogen catabolite repression-sensitive genes that encode enzyme and transport proteins, the promoters of the GAT1, DAL80, and DEH1 genes all contain multiple GATA sequences as well. These GATA sequences are the binding sites of the GATA family transcription factors and are hypothesized to mediate their autogenous and cross regulation. Here we show, using DAL80 fused to the carbon-regulated GAL1,10 or copper-regulated CUP1 promoter, that GAT1 expression is inversely regulated by the level of DAL80 expression, i.e., as DAL80 expression increases, GAT1 expression decreases. The amount of DAL80 expression also dictates the level at which DAL3, a gene activated almost exclusively by Gln3p, is transcribed. Gat1p was found to partially substitute for Gln3p in transcription. These data support the contention that regulation of GATA-factor gene expression is tightly and dynamically coupled. Finally, we suggest that the complicated regulatory circuit in which the GATA family transcription factors participate is probably most beneficial as cells make the transition from excess to limited nitrogen availability.
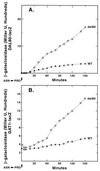
Figures




Similar articles
-
Transmitting the signal of excess nitrogen in Saccharomyces cerevisiae from the Tor proteins to the GATA factors: connecting the dots.FEMS Microbiol Rev. 2002 Aug;26(3):223-38. doi: 10.1111/j.1574-6976.2002.tb00612.x. FEMS Microbiol Rev. 2002. PMID: 12165425 Free PMC article. Review.
-
Cross regulation of four GATA factors that control nitrogen catabolic gene expression in Saccharomyces cerevisiae.J Bacteriol. 1997 Jun;179(11):3416-29. doi: 10.1128/jb.179.11.3416-3429.1997. J Bacteriol. 1997. PMID: 9171383 Free PMC article.
-
Nitrogen catabolite repression of DAL80 expression depends on the relative levels of Gat1p and Ure2p production in Saccharomyces cerevisiae.J Biol Chem. 2000 May 12;275(19):14408-14. doi: 10.1074/jbc.275.19.14408. J Biol Chem. 2000. PMID: 10799523 Free PMC article.
-
Expression of the DAL80 gene, whose product is homologous to the GATA factors and is a negative regulator of multiple nitrogen catabolic genes in Saccharomyces cerevisiae, is sensitive to nitrogen catabolite repression.Mol Cell Biol. 1991 Dec;11(12):6205-15. doi: 10.1128/mcb.11.12.6205-6215.1991. Mol Cell Biol. 1991. PMID: 1944286 Free PMC article.
-
Nitrogen catabolite repression in Saccharomyces cerevisiae.Mol Biotechnol. 1999 Aug;12(1):35-73. doi: 10.1385/MB:12:1:35. Mol Biotechnol. 1999. PMID: 10554772 Review.
Cited by
-
Nuclear Gln3 Import Is Regulated by Nitrogen Catabolite Repression Whereas Export Is Specifically Regulated by Glutamine.Genetics. 2015 Nov;201(3):989-1016. doi: 10.1534/genetics.115.177725. Epub 2015 Sep 2. Genetics. 2015. PMID: 26333687 Free PMC article.
-
The yeast GATA factor Gat1 occupies a central position in nitrogen catabolite repression-sensitive gene activation.Mol Cell Biol. 2009 Jul;29(13):3803-15. doi: 10.1128/MCB.00399-09. Epub 2009 Apr 20. Mol Cell Biol. 2009. PMID: 19380492 Free PMC article.
-
Nitrogen catabolite repression-sensitive transcription as a readout of Tor pathway regulation: the genetic background, reporter gene and GATA factor assayed determine the outcomes.Genetics. 2009 Mar;181(3):861-74. doi: 10.1534/genetics.108.099051. Epub 2008 Dec 22. Genetics. 2009. PMID: 19104072 Free PMC article.
-
An evolutionary and functional assessment of regulatory network motifs.Genome Biol. 2005;6(4):R35. doi: 10.1186/gb-2005-6-4-r35. Epub 2005 Mar 24. Genome Biol. 2005. PMID: 15833122 Free PMC article.
-
Transmitting the signal of excess nitrogen in Saccharomyces cerevisiae from the Tor proteins to the GATA factors: connecting the dots.FEMS Microbiol Rev. 2002 Aug;26(3):223-38. doi: 10.1111/j.1574-6976.2002.tb00612.x. FEMS Microbiol Rev. 2002. PMID: 12165425 Free PMC article. Review.
References
-
- Beck T, Hall M N. The TOR signaling pathway controls nuclear localization of nutrient-regulated transcription factors. Nature. 1999;402:689–692. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

