Essential regions of the tRNA primer required for HIV-1 infectivity
- PMID: 11095691
- PMCID: PMC115161
- DOI: 10.1093/nar/28.23.4783
Essential regions of the tRNA primer required for HIV-1 infectivity
Abstract
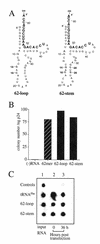
Human immunodeficiency virus (HIV), like all retroviruses, requires a cellular tRNA as a primer for initiation of reverse transcription. In a previous study, we demonstrated that an HIV-1 with a primer binding site complementary to yeast tRNA(Phe) (psHIV-Phe) was not infectious unless yeast tRNA(Phe) was supplied in trans. This unique in vivo complementation system has now been used to define the elements of the tRNA required for HIV-1 replication. Mutant tRNA(Phe) with deletions in TPsiC stem-loop, anticodon stem-loop or D stem-loop of the tRNA were generated and assessed for the capacity to rescue psHIV-Phe. Mutant tRNA(Phe) with disrupted TPsiC stem-loop did not rescue psHIV-Phe. In contrast, a mutant tRNA(Phe) without the D stem-loop was fully functional for the rescue. The tRNA anticodon stem-loop region was found to be important for efficient complementation. The results of our studies demonstrate for the first time the importance of specific structural and sequence elements of the tRNA primer for HIV-1 reverse transcription and define new targets for interruption of HIV-1 replication.
Figures




Similar articles
-
Identification of critical elements in the tRNA acceptor stem and T(Psi)C loop necessary for human immunodeficiency virus type 1 infectivity.J Virol. 2001 May;75(10):4902-6. doi: 10.1128/JVI.75.10.4902-4906.2001. J Virol. 2001. PMID: 11312362 Free PMC article.
-
Yeast tRNA(Phe) expressed in human cells can be selected by HIV-1 for use as a reverse transcription primer.Virology. 2003 Sep 1;313(2):354-63. doi: 10.1016/s0042-6822(03)00243-5. Virology. 2003. PMID: 12954204
-
Selection of retroviral reverse transcription primer is coordinated with tRNA biogenesis.J Virol. 2003 Aug;77(16):8695-701. doi: 10.1128/jvi.77.16.8695-8701.2003. J Virol. 2003. PMID: 12885888 Free PMC article.
-
HIV-1 reverse transcription initiation: a potential target for novel antivirals?Virus Res. 2008 Jun;134(1-2):4-18. doi: 10.1016/j.virusres.2007.12.009. Epub 2008 Feb 6. Virus Res. 2008. PMID: 18255184 Review.
-
Structural bases of the annealing of primer tRNA(3Lys) to the HIV-1 viral RNA.Curr HIV Res. 2005 Apr;3(2):147-56. doi: 10.2174/1570162053506919. Curr HIV Res. 2005. PMID: 15853720 Review.
Cited by
-
Structural elements of the tRNA TPsiC loop critical for nucleocytoplasmic transport are important for human immunodeficiency virus type 1 primer selection.J Virol. 2005 May;79(10):6532-9. doi: 10.1128/JVI.79.10.6532-6539.2005. J Virol. 2005. PMID: 15858038 Free PMC article.
-
Identification of specific HIV-1 reverse transcriptase contacts to the viral RNA:tRNA complex by mass spectrometry and a primary amine selective reagent.Proc Natl Acad Sci U S A. 2002 Dec 10;99(25):15988-93. doi: 10.1073/pnas.252550199. Epub 2002 Dec 2. Proc Natl Acad Sci U S A. 2002. PMID: 12461175 Free PMC article.
-
Nucleotides within the anticodon stem are important for optimal use of tRNA(Lys,3) as the primer for HIV-1 reverse transcription.Virology. 2007 Jul 20;364(1):169-77. doi: 10.1016/j.virol.2007.02.010. Epub 2007 Mar 21. Virology. 2007. PMID: 17368706 Free PMC article.
-
Mutations in the TPsiC loop of E. coli tRNALys,3 have varied effects on in trans complementation of HIV-1 replication.Virol J. 2007 Jan 11;4:5. doi: 10.1186/1743-422X-4-5. Virol J. 2007. PMID: 17217532 Free PMC article.
-
Influence of L-lysine amino acid on the HIV-1 RNA replication in vitro.Antivir Chem Chemother. 2015 Feb;24(1):39-46. doi: 10.1177/2040206614566582. Antivir Chem Chemother. 2015. PMID: 26149265 Free PMC article.
References
-
- Kang S.M., Wakefield,J.K. and Morrow,C.D. (1996) Virology, 222, 401–414. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

