The repertoire for pattern recognition of pathogens by the innate immune system is defined by cooperation between toll-like receptors
- PMID: 11095740
- PMCID: PMC17650
- DOI: 10.1073/pnas.250476497
The repertoire for pattern recognition of pathogens by the innate immune system is defined by cooperation between toll-like receptors
Abstract
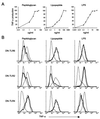
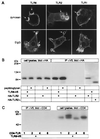
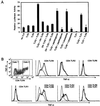
Toll-like receptors (TLRs) have been shown to participate in the recognition of pathogens by the innate immune system, but it is not clear how a restricted family of receptors has the capacity to recognize the wide spectrum of TLR stimuli known to exist. We report here that two members of the TLR family, TLR2 and TLR6, together coordinate macrophage activation by Gram-positive bacteria and the yeast cell-wall particle, zymosan. TLR6 and TLR2 both are recruited to the macrophage phagosome, where they recognize peptidoglycan, a Gram-positive pathogen component. By contrast, TLR2 recognizes another component, bacterial lipopeptide, without TLR6. The requirement for TLR cooperation is supported by the finding that TLR2 needs a partner to activate tumor necrosis factor-alpha production in macrophages. Dimerization of the cytoplasmic domain of TLR2 does not induce tumor necrosis factor-alpha production in macrophages, whereas similar dimerization of the TLR4 cytoplasmic domain does. We show that the cytoplasmic domain of TLR2 can form functional pairs with TLR6 or TLR1, and this interaction leads to cytokine induction. Thus, the cytoplasmic tails of TLRs are not functionally equivalent, with certain TLRs requiring assembly into heteromeric complexes, whereas others are active as homomeric complexes. Finally, we show that TLR6, TLR2, and TLR1 are recruited to macrophage phagosomes that contain IgG-coated erythrocytes that do not display microbial components. The data suggest that TLRs sample the contents of the phagosome independent of the nature of the contents, and can establish a combinatorial repertoire to discriminate among the large number of pathogen-associated molecular patterns found in nature.
Figures

References
-
- Janeway C A., Jr Cold Spring Harbor Symp Quant Biol. 1989;54:1–13.
-
- Janeway C A., Jr Immunol Today. 1992;13:11–16. - PubMed
-
- Underhill D M, Ozinsky A, Hajjar A M, Stevens A, Wilson C B, Bassetti M, Aderem A. Nature (London) 1999;401:811–815. - PubMed
-
- Takeuchi O, Hoshino K, Kawai T, Sanjo H, Takada H, Ogawa T, Takeda K, Akira S. Immunity. 1999;11:443–451. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

