The sodium/proton exchanger Nhx1p is required for endosomal protein trafficking in the yeast Saccharomyces cerevisiae
- PMID: 11102523
- PMCID: PMC15072
- DOI: 10.1091/mbc.11.12.4277
The sodium/proton exchanger Nhx1p is required for endosomal protein trafficking in the yeast Saccharomyces cerevisiae
Abstract
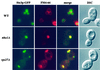
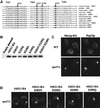
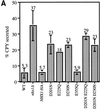
We show that the vacuolar protein sorting gene VPS44 is identical to NHX1, a gene that encodes a sodium/proton exchanger. The Saccharomyces cerevisiae protein Nhx1p shows high homology to mammalian sodium/proton exchangers of the NHE family. Nhx1p is thought to transport sodium ions into the prevacuole compartment in exchange for protons. Pulse-chase experiments show that approximately 35% of the newly synthesized soluble vacuolar protein carboxypeptidase Y is missorted in nhx1 delta cells, and is secreted from the cell. nhx1 delta cells accumulate late Golgi, prevacuole, and lysosome markers in an aberrant structure next to the vacuole, and late Golgi proteins are proteolytically cleaved more rapidly than in wild-type cells. Our results show that efficient transport out of the prevacuolar compartment requires Nhx1p, and that nhx1 delta cells exhibit phenotypes characteristic of the "class E" group of vps mutants. In addition, we show that Nhx1p is required for protein trafficking even in the absence of the vacuolar ATPase. Our analysis of Nhx1p provides the first evidence that a sodium/proton exchange protein is important for correct protein sorting, and that intraorganellar ion balance may be important for endosomal function in yeast.
Figures

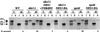







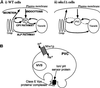
Similar articles
-
The endosomal Na(+)/H(+) exchanger contributes to multivesicular body formation by regulating the recruitment of ESCRT-0 Vps27p to the endosomal membrane.J Biol Chem. 2011 Oct 28;286(43):37625-38. doi: 10.1074/jbc.M111.260612. Epub 2011 Sep 5. J Biol Chem. 2011. PMID: 21896492 Free PMC article.
-
Multilamellar endosome-like compartment accumulates in the yeast vps28 vacuolar protein sorting mutant.Mol Biol Cell. 1996 Jun;7(6):985-99. doi: 10.1091/mbc.7.6.985. Mol Biol Cell. 1996. PMID: 8817003 Free PMC article.
-
Localization and functional requirement of yeast Na+/H+ exchanger, Nhx1p, in the endocytic and protein recycling pathway.Biochim Biophys Acta. 2012 Feb;1823(2):534-43. doi: 10.1016/j.bbamcr.2011.12.004. Epub 2011 Dec 23. Biochim Biophys Acta. 2012. PMID: 22210050
-
Protein transport from the late Golgi to the vacuole in the yeast Saccharomyces cerevisiae.Biochim Biophys Acta. 2005 Jul 10;1744(3):438-54. doi: 10.1016/j.bbamcr.2005.04.004. Biochim Biophys Acta. 2005. PMID: 15913810 Review.
-
V-ATPase, ScNhx1p and yeast vacuole fusion.J Genet Genomics. 2012 Apr 20;39(4):167-71. doi: 10.1016/j.jgg.2012.02.001. Epub 2012 Feb 10. J Genet Genomics. 2012. PMID: 22546538 Review.
Cited by
-
Different evolutionary histories of two cation/proton exchanger gene families in plants.BMC Plant Biol. 2013 Jul 4;13:97. doi: 10.1186/1471-2229-13-97. BMC Plant Biol. 2013. PMID: 23822194 Free PMC article.
-
The endosomal Na(+)/H(+) exchanger contributes to multivesicular body formation by regulating the recruitment of ESCRT-0 Vps27p to the endosomal membrane.J Biol Chem. 2011 Oct 28;286(43):37625-38. doi: 10.1074/jbc.M111.260612. Epub 2011 Sep 5. J Biol Chem. 2011. PMID: 21896492 Free PMC article.
-
AtNHX5 and AtNHX6 Are Required for the Subcellular Localization of the SNARE Complex That Mediates the Trafficking of Seed Storage Proteins in Arabidopsis.PLoS One. 2016 Mar 17;11(3):e0151658. doi: 10.1371/journal.pone.0151658. eCollection 2016. PLoS One. 2016. PMID: 26986836 Free PMC article.
-
Expression and integrated network analyses revealed functional divergence of NHX-type Na+/H+ exchanger genes in poplar.Sci Rep. 2017 Jun 1;7(1):2607. doi: 10.1038/s41598-017-02894-8. Sci Rep. 2017. PMID: 28572621 Free PMC article.
-
The Na+(K+)/H+ exchanger Nhx1 controls multivesicular body-vacuolar lysosome fusion.Mol Biol Cell. 2018 Feb 1;29(3):317-325. doi: 10.1091/mbc.E17-08-0496. Epub 2017 Dec 6. Mol Biol Cell. 2018. PMID: 29212874 Free PMC article.
References
-
- Apse MP, Aharon GS, Snedden WA, Blumwald E. Salt tolerance conferred by overexpression of a vacuolar Na+/H+ antiport in Arabidopsis. Science. 1999;285:1256–1258. - PubMed
-
- Babst M, Odorizzi G, Estepa EJ, Emr SD. Mammalian tumor susceptibility gene 101 (TSG101) and the yeast homologue, Vps23p, both function in late endosomal trafficking. Traffic. 2000;1:248–258. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

