Coupling of flagellar gene expression to flagellar assembly in Salmonella enterica serovar typhimurium and Escherichia coli
- PMID: 11104815
- PMCID: PMC99010
- DOI: 10.1128/MMBR.64.4.694-708.2000
Coupling of flagellar gene expression to flagellar assembly in Salmonella enterica serovar typhimurium and Escherichia coli
Abstract
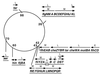
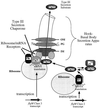
How do organisms assess the degree of completion of a large structure, especially an extracellular structure such as a flagellum? Bacteria can do this. Mutants that lack key components needed early in assembly fail to express proteins that would normally be added at later assembly stages. In some cases, the regulatory circuitry is able to sense completion of structures beyond the cell surface, such as completion of the external hook structure. In Salmonella and Escherichia coli, regulation occurs at both transcriptional and posttranscriptional levels. One transcriptional regulatory mechanism involves a regulatory protein, FlgM, that escapes from the cell (and thus can no longer act) through a complete flagellum and is held inside when the structure has not reached a later stage of completion. FlgM prevents late flagellar gene transcription by binding the flagellum-specific transcription factor sigma(28). FlgM is itself regulated in response to the assembly of an incomplete flagellum known as the hook-basal body intermediate structure. Upon completion of the hook-basal body structure, FlgM is exported through this structure out of the cell. Inhibition of sigma(28)-dependent transcription is relieved, and genes required for the later assembly stages are expressed, allowing completion of the flagellar organelle. Distinct posttranscriptional regulatory mechanisms occur in response to assembly of the flagellar type III secretion apparatus and of ring structures in the peptidoglycan and lipopolysaccharide layers. The entire flagellar regulatory pathway is regulated in response to environmental cues. Cell cycle control and flagellar development are codependent. We discuss how all these levels of regulation ensure efficient assembly of the flagellum in response to environmental stimuli.
Figures



References
-
- Adler J, Templeton B. The effect of environmental conditions on the motility of Escherichia coli. J Gen Microbiol. 1967;46:175–184. - PubMed
-
- Aizawa S-I. Flagellar assembly in Salmonella typhimurium. Mol Microbiol. 1996;20:1–4. - PubMed
-
- Aizawa S-I, Kubori T. Bacterial flagellation and cell division. Genes Cells. 1998;3:625–634. - PubMed
-
- Aizawa S-I, Vonderviszt F, Ishima R, Akasaka K. Termini of Salmonella flagellin are disordered and become organized upon polymerization into flagellar filament. J Mol Biol. 1990;211:673–677. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

