The gaf fimbrial gene cluster of Escherichia coli expresses a full-size and a truncated soluble adhesin protein
- PMID: 11133944
- PMCID: PMC94906
- DOI: 10.1128/JB.183.2.512-519.2001
The gaf fimbrial gene cluster of Escherichia coli expresses a full-size and a truncated soluble adhesin protein
Abstract
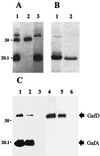
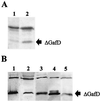
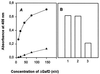
The GafD lectin of the G (F17) fimbriae of diarrhea-associated Escherichia coli was overexpressed and purified from the periplasm of E. coli by affinity chromatography on GlcNAc-agarose. The predicted mature GafD peptide comprises 321 amino acids, but the predominant form of GafD recovered from the periplasm was 19,092 Da in size and corresponded to the 178 N-terminal amino acid residues, as judged by mass spectrometry and amino acid sequencing, and was named DeltaGafD. Expression of gafD from the cloned gaf gene cluster in DegP-, Lon-, and OmpT-deficient recombinant strains did not significantly decrease the formation of DeltaGafD. The peptide was also detected in the periplasm of the wild-type E. coli strain from which the gaf gene cluster originally was cloned. We expressed gafD fragments encoding C-terminally truncated peptides. Peptides GafD1-252, GafD1-224, GafD1-189, and the GafD1-178, isolated from the periplasm by affinity chromatography, had apparent sizes closely similar to that of DeltaGafD. Only trace amounts of truncated forms with expected molecular sizes were detected in spheroplasts. In contrast, the shorter GafD1-157 peptide was detected in spheroplasts but not in the periplasm, indicating that it was poorly translocated or was degraded by periplasmic proteases. Pulse-chase assays using gafD indicated that DeltaGafD was processed from GafD and is not a primary translation product. The DeltaGafD peptide was soluble by biochemical criteria and exhibited specific binding to GlcNAc-agarose. Inhibition assays with mono- and oligosaccharides gave a similar inhibition pattern in the hemagglutination by the G-fimbria-expressing recombinant E. coli strain and in the binding of [(14)C]DeltaGafD to GlcNAc-agarose. DeltaGafD bound specifically to laminin, a previously described tissue target for the G fimbria. Our results show that a soluble, protease-resistant subdomain of GafD exhibits receptor-binding specificity similar to that for intact G fimbriae and that it is formed when gafD is expressed alone or from the gaf gene cluster.
Figures

Similar articles
-
The Escherichia coli G-fimbrial lectin protein participates both in fimbrial biogenesis and in recognition of the receptor N-acetyl-D-glucosamine.J Bacteriol. 1995 Mar;177(6):1477-84. doi: 10.1128/jb.177.6.1477-1484.1995. J Bacteriol. 1995. PMID: 7883703 Free PMC article.
-
The GafD protein of the G (F17) fimbrial complex confers adhesiveness of Escherichia coli to laminin.Infect Immun. 1996 Jul;64(7):2857-60. doi: 10.1128/iai.64.7.2857-2860.1996. Infect Immun. 1996. PMID: 8698525 Free PMC article.
-
Characterization of 20K fimbria, a new adhesin of septicemic and diarrhea-associated Escherichia coli strains, that belongs to a family of adhesins with N-acetyl-D-glucosamine recognition.Infect Immun. 1996 Jan;64(1):332-42. doi: 10.1128/iai.64.1.332-342.1996. Infect Immun. 1996. PMID: 8557360 Free PMC article.
-
AFA and F17 adhesins produced by pathogenic Escherichia coli strains in domestic animals.Vet Res. 1999 Mar-Jun;30(2-3):317-42. Vet Res. 1999. PMID: 10367361 Review.
-
Molecular structure of adhesin domains in Escherichia coli fimbriae.Int J Med Microbiol. 2005 Oct;295(6-7):479-86. doi: 10.1016/j.ijmm.2005.06.010. Int J Med Microbiol. 2005. PMID: 16238022 Review.
Cited by
-
A newly identified leptospiral adhesin mediates attachment to laminin.Infect Immun. 2006 Nov;74(11):6356-64. doi: 10.1128/IAI.00460-06. Epub 2006 Sep 5. Infect Immun. 2006. PMID: 16954400 Free PMC article.
-
A simple strategy for the creation of a recombinant lectin microarray.Mol Biosyst. 2008 Jun;4(6):654-62. doi: 10.1039/b800725j. Epub 2008 Apr 14. Mol Biosyst. 2008. PMID: 18493664 Free PMC article.
-
Identification of a Treponema pallidum laminin-binding protein.Infect Immun. 2003 May;71(5):2525-33. doi: 10.1128/IAI.71.5.2525-2533.2003. Infect Immun. 2003. PMID: 12704124 Free PMC article.
-
Targeted in vivo O-GlcNAc sensors reveal discrete compartment-specific dynamics during signal transduction.J Biol Chem. 2011 Feb 25;286(8):6650-8. doi: 10.1074/jbc.M110.191627. Epub 2010 Dec 7. J Biol Chem. 2011. PMID: 21138847 Free PMC article.
-
Mapping the binding domain of the F18 fimbrial adhesin.Infect Immun. 2003 Apr;71(4):2163-72. doi: 10.1128/IAI.71.4.2163-2182.2003. Infect Immun. 2003. PMID: 12654838 Free PMC article.
References
-
- Arumugham R G, Hsieh T C-Y, Tanzer M L, Laine R A. Structures of the asparagine-linked sugar chains of laminin. Biochim Biophys Acta. 1986;883:112–126. - PubMed
-
- Bakker D, Vader C E M, Roosendaal B, Mooi F R, Oudega B, de Graaf F K. Structure and function of periplasmic chaperone-like proteins involved in the biosynthesis of K88 and K99 fimbriae in enterotoxigenic Escherichia coli. Mol Microbiol. 1991;5:867–886. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

