Heterogeneous geographic patterns of nucleotide sequence diversity between two alcohol dehydrogenase genes in wild barley (Hordeum vulgare subspecies spontaneum)
- PMID: 11149938
- PMCID: PMC14621
- DOI: 10.1073/pnas.98.2.531
Heterogeneous geographic patterns of nucleotide sequence diversity between two alcohol dehydrogenase genes in wild barley (Hordeum vulgare subspecies spontaneum)
Abstract
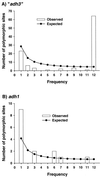
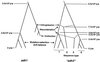
Patterns of nucleotide sequence diversity in the predominantly self-fertilizing species Hordeum vulgare subspecies spontaneum (wild barley) are compared between the putative alcohol dehydrogenase 3 locus (denoted "adh3") and alcohol dehydrogenase 1 (adh1), two related but unlinked loci. The data consist of a sequence sample of 1,873 bp of "adh3" drawn from 25 accessions that span the species range. There were 104 polymorphic sites in the sequenced region of "adh3." The data reveal a strong geographic pattern of diversity at "adh3" despite geographic uniformity at adh1. Moreover, levels of nucleotide sequence diversity differ by nearly an order of magnitude between the two loci. Genealogical analysis resolved two distinct clusters of "adh3" alleles (dimorphic sequence types) that coalesce roughly 3 million years ago. One type consists of accessions from the Middle East, and the other consists of accessions predominantly from the Near East. The two "adh3" sequence types are characterized by a high level of differentiation between clusters ( approximately 2.2%), which induces an overall excess of intermediate frequency variants in the pooled sample. Finally, there is evidence of intralocus recombination in the "adh3" data, despite the high level of self-fertilization characteristic of wild barley.
Figures



References
-
- Kingman J F C. Stochastic Processes: Formalism. Appl Proc Winter Sch. 1982;13:235–248.
-
- Hudson R. Oxford Surv Evol Biol. 1990;7:1–44.
-
- Clegg M T. Heredity. 1997;88:1–7. - PubMed
-
- von Bothmer R, Jacobsen N, Baden C, Jorgensen R B, Linde-Laursen I. An Ecogeographical Study of the Genus Hordeum. 2nd Ed. Rome: International Plant Genetic Resources Institute; 1995.
-
- Brown A H D, Zohary D, Nevo E. Heredity. 1978;41:49–62.
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources

