Novel bacterial lineages at the (sub)division level as detected by signature nucleotide-targeted recovery of 16S rRNA genes from bulk soil and rice roots of flooded rice microcosms
- PMID: 11157225
- PMCID: PMC92629
- DOI: 10.1128/AEM.67.2.623-631.2001
Novel bacterial lineages at the (sub)division level as detected by signature nucleotide-targeted recovery of 16S rRNA genes from bulk soil and rice roots of flooded rice microcosms
Abstract
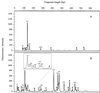
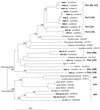
Using a newly developed 16S rRNA gene (rDNA)-targeted PCR assay with proposed group specificity for planctomycetes, we examined anoxic bulk soil of flooded rice microcosms for the presence of novel planctomycete-like diversity. For comparison, oxic rice roots were included as an additional sample in this investigation. The bacterial diversity detectable by this PCR assay was assessed by using a combined approach that included terminal restriction fragment length polymorphism (T-RFLP) analysis and comparative sequence analysis of cloned 16S rDNA. T-RFLP fingerprint patterns generated from rice roots contained 12 distinct terminal restriction fragments (T-RFs). In contrast, the T-RFLP fingerprint patterns obtained from the anoxic bulk soil contained 33 distinct T-RFs, a clearly higher level of complexity. A survey of 176 bulk soil 16S rDNA clone sequences permitted correlation of 20 T-RFs with phylogenetic information. The other 13 T-RFs remained unidentified. The predominant T-RFs obtained from rice roots could be assigned to members of the genus Pirellula within the Planctomycetales, while most of the T-RFs obtained from the bulk soil corresponded to novel lines of bacterial descent. Using a level of 16S rDNA sequence dissimilarity to cultured microorganisms of approximately 20% as a threshold value, we detected 11 distinct bacterial lineages for which pure-culture representatives are not known. Four of these lineages could be assigned to the order Planctomycetales, while one lineage was affiliated with the division Verrucomicrobia and one lineage was affiliated with the spirochetes. The other five lineages either could not be assigned to any of the main lines of bacterial descent or clearly expanded the known diversity of division level lineages WS3 and OP3. Our results indicate the presence of bacterial diversity at a subdivision and/or division level that has not been detected previously by the so-called universal 16S rDNA PCR assays.
Figures

Similar articles
-
Detection of methanotroph diversity on roots of submerged rice plants by molecular retrieval of pmoA, mmoX, mxaF, and 16S rRNA and ribosomal DNA, including pmoA-based terminal restriction fragment length polymorphism profiling.Appl Environ Microbiol. 2001 Sep;67(9):4177-85. doi: 10.1128/AEM.67.9.4177-4185.2001. Appl Environ Microbiol. 2001. PMID: 11526021 Free PMC article.
-
Spatial changes in the bacterial community structure along a vertical oxygen gradient in flooded paddy soil cores.Appl Environ Microbiol. 2000 Feb;66(2):754-62. doi: 10.1128/AEM.66.2.754-762.2000. Appl Environ Microbiol. 2000. PMID: 10653747 Free PMC article.
-
Comparative phylogenetic assignment of environmental sequences of genes encoding 16S rRNA and numerically abundant culturable bacteria from an anoxic rice paddy soil.Appl Environ Microbiol. 1999 Nov;65(11):5050-8. doi: 10.1128/AEM.65.11.5050-5058.1999. Appl Environ Microbiol. 1999. PMID: 10543822 Free PMC article.
-
Impact of flooding on soil bacterial communities associated with poplar (Populus sp.) trees.FEMS Microbiol Ecol. 2005 Aug 1;53(3):401-15. doi: 10.1016/j.femsec.2005.01.009. FEMS Microbiol Ecol. 2005. PMID: 16329959
-
Identifying the dominant soil bacterial taxa in libraries of 16S rRNA and 16S rRNA genes.Appl Environ Microbiol. 2006 Mar;72(3):1719-28. doi: 10.1128/AEM.72.3.1719-1728.2006. Appl Environ Microbiol. 2006. PMID: 16517615 Free PMC article. Review. No abstract available.
Cited by
-
Gene discovery within the planctomycete division of the domain Bacteria using sequence tags from genomic DNA libraries.Genome Biol. 2002;3(6):RESEARCH0031. doi: 10.1186/gb-2002-3-6-research0031. Epub 2002 May 30. Genome Biol. 2002. PMID: 12093378 Free PMC article.
-
Phylum- and class-specific PCR primers for general microbial community analysis.Appl Environ Microbiol. 2005 Oct;71(10):6193-8. doi: 10.1128/AEM.71.10.6193-6198.2005. Appl Environ Microbiol. 2005. PMID: 16204538 Free PMC article.
-
Characterization of an autotrophic nitrogen-removing biofilm from a highly loaded lab-scale rotating biological contactor.Appl Environ Microbiol. 2003 Jun;69(6):3626-35. doi: 10.1128/AEM.69.6.3626-3635.2003. Appl Environ Microbiol. 2003. PMID: 12788771 Free PMC article.
-
Spatiotemporal analysis of bacterial diversity in sediments of Sundarbans using parallel 16S rRNA gene tag sequencing.Microb Ecol. 2015 Apr;69(3):500-11. doi: 10.1007/s00248-014-0498-y. Epub 2014 Sep 26. Microb Ecol. 2015. PMID: 25256302
-
Energy and carbon metabolisms in a deep terrestrial subsurface fluid microbial community.ISME J. 2017 Oct;11(10):2319-2333. doi: 10.1038/ismej.2017.94. Epub 2017 Jun 23. ISME J. 2017. PMID: 28644444 Free PMC article.
References
-
- Defosse D L, Johnson R C, Paster B J, Dewhirst F E, Fraser G J. Brevinema andersonii gen. nov., sp. nov., an infectious spirochete isolated from the short-tailed shrew (Blarina brevicauda) and the white-footed mouse (Peromyscus leucopus) Int J Syst Bacteriol. 1995;45:78–84. - PubMed
-
- DeLong E F, Franks D G, Alldredge A L. Phylogenetic diversity of aggregate-attached vs. free-living marine bacterial assemblages. Limnol Oceanogr. 1993;38:924–934.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

