Dissection of central carbon metabolism of hemoglobin-expressing Escherichia coli by 13C nuclear magnetic resonance flux distribution analysis in microaerobic bioprocesses
- PMID: 11157231
- PMCID: PMC92635
- DOI: 10.1128/AEM.67.2.680-687.2001
Dissection of central carbon metabolism of hemoglobin-expressing Escherichia coli by 13C nuclear magnetic resonance flux distribution analysis in microaerobic bioprocesses
Abstract
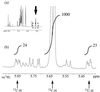
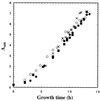
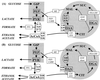
Escherichia coli MG1655 cells expressing Vitreoscilla hemoglobin (VHb), Alcaligenes eutrophus flavohemoprotein (FHP), the N-terminal hemoglobin domain of FHP (FHPg), and a fusion protein which comprises VHb and the A. eutrophus C-terminal reductase domain (VHb-Red) were grown in a microaerobic bioreactor to study the effects of low oxygen concentrations on the central carbon metabolism, using fractional (13)C-labeling of the proteinogenic amino acids and two-dimensional [(13)C, (1)H]-correlation nuclear magnetic resonance (NMR) spectroscopy. The NMR data revealed differences in the intracellular carbon fluxes between E. coli cells expressing either VHb or VHb-Red and cells expressing A. eutrophus FHP or the truncated heme domain (FHPg). E. coli MG1655 cells expressing either VHb or VHb-Red were found to function with a branched tricarboxylic acid (TCA) cycle. Furthermore, cellular demands for ATP and reduction equivalents in VHb- and VHb-Red-expressing cells were met by an increased flux through glycolysis. In contrast, in E. coli cells expressing A. eutrophus hemeproteins, the TCA cycle is running cyclically, indicating a shift towards a more aerobic regulation. Consistently, E. coli cells displaying FHP and FHPg activity showed lower production of the typical anaerobic by-products formate, acetate, and D-lactate. The implications of these observations for biotechnological applications are discussed.
Figures
Similar articles
-
Expression of Alcaligenes eutrophus flavohemoprotein and engineered Vitreoscilla hemoglobin-reductase fusion protein for improved hypoxic growth of Escherichia coli.Appl Environ Microbiol. 2000 Jan;66(1):98-104. doi: 10.1128/AEM.66.1.98-104.2000. Appl Environ Microbiol. 2000. PMID: 10618209 Free PMC article.
-
Novel hemoglobins to enhance microaerobic growth and substrate utilization in Escherichia coli.Biotechnol Prog. 2001 Sep-Oct;17(5):798-808. doi: 10.1021/bp010091j. Biotechnol Prog. 2001. PMID: 11587567
-
Error-prone PCR of Vitreoscilla hemoglobin (VHb) to support the growth of microaerobic Escherichia coli.Biotechnol Bioeng. 2000 Nov 20;70(4):446-55. Biotechnol Bioeng. 2000. PMID: 11005927
-
The single-domain globin of Vitreoscilla: augmentation of aerobic metabolism for biotechnological applications.Adv Microb Physiol. 2011;58:81-139. doi: 10.1016/B978-0-12-381043-4.00003-9. Adv Microb Physiol. 2011. PMID: 21722792 Review.
-
Bacterial hemoglobins and flavohemoglobins: versatile proteins and their impact on microbiology and biotechnology.FEMS Microbiol Rev. 2003 Oct;27(4):525-45. doi: 10.1016/S0168-6445(03)00056-1. FEMS Microbiol Rev. 2003. PMID: 14550944 Review.
Cited by
-
The Biochemistry of Vitreoscilla hemoglobin.Comput Struct Biotechnol J. 2012 Oct 29;3:e201210002. doi: 10.5936/csbj.201210002. eCollection 2012. Comput Struct Biotechnol J. 2012. PMID: 24688662 Free PMC article. Review.
-
Recent Advances in the Physicochemical Properties and Biotechnological Application of Vitreoscilla Hemoglobin.Microorganisms. 2021 Jul 7;9(7):1455. doi: 10.3390/microorganisms9071455. Microorganisms. 2021. PMID: 34361891 Free PMC article. Review.
-
Physiological Response of Escherichia coli W3110 and BL21 to the Aerobic Expression of Vitreoscilla Hemoglobin.J Microbiol Biotechnol. 2020 Oct 28;30(10):1592-1596. doi: 10.4014/jmb.2004.04030. J Microbiol Biotechnol. 2020. PMID: 32699196 Free PMC article.
-
Bacterial hemoglobins and flavohemoglobins for alleviation of nitrosative stress in Escherichia coli.Appl Environ Microbiol. 2002 Oct;68(10):4835-40. doi: 10.1128/AEM.68.10.4835-4840.2002. Appl Environ Microbiol. 2002. PMID: 12324328 Free PMC article.
References
-
- Amarasingham C R, Davis B C. Regulation of δ-ketoglutarate dehydrogenase formation in Escherichia coli. J Biol Chem. 1965;240:3664–3668. - PubMed
-
- Babul J, Clifton D, Kretschmer M, Fraenkel D G. Glucose metabolism in Escherichia coli and the effect of increased amount of aldolase. Biochemistry. 1993;32:4685–4692. - PubMed
-
- Bailey J E, Sburlati A, Hatzimanikatis V, Lee K, Renner W A, Tsai P S. Inverse metabolic engineering: a strategy for directed genetic engineering of useful phenotypes. Biotechnol Bioeng. 1996;52:109–121. - PubMed
-
- Bax A, Pochapsky S. Optimized recording of heteronuclear multidimensional NMR spectra using pulsed field gradients. J Magn Reson. 1992;99:638–643.
-
- Bergmeyer J, Gassl M. Metabolites 1: carbohydrates. WeinheimWeinheim, Germany: VCH; 1984.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

