Identification of Mur, an atypical peptidoglycan hydrolase derived from Leuconostoc citreum
- PMID: 11157255
- PMCID: PMC92659
- DOI: 10.1128/AEM.67.2.858-864.2001
Identification of Mur, an atypical peptidoglycan hydrolase derived from Leuconostoc citreum
Abstract
A gene encoding a protein homologous to known bacterial N-acetyl-muramidases has been cloned from Leuconostoc citreum by a PCR-based approach. The encoded protein, Mur, consists of 209 amino acid residues with a calculated molecular mass of 23,821 Da including a 31-amino-acid putative signal peptide. In contrast to most of the other known peptidoglycan hydrolases, L. citreum Mur protein does not contain amino acid repeats involved in cell wall binding. The purified L. citreum Mur protein was shown to exhibit peptidoglycan-hydrolyzing activity by renaturing sodium dodecyl sulfate-polyacrylamide gel electrophoresis. An active chimeric protein was constructed by fusion of L. citreum Mur to the C-terminal repeat-containing domain (cA) of AcmA, the major autolysin of Lactococcus lactis. Expression of the Mur-cA fusion protein was able to complement an acmA mutation in L. lactis; normal cell separation after cell division was restored by Mur-cA expression.
Figures


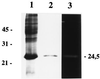
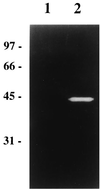

Similar articles
-
Analysis of the peptidoglycan hydrolase complement of Lactococcus lactis: identification of a third N-acetylglucosaminidase, AcmC.Appl Environ Microbiol. 2004 Jun;70(6):3493-9. doi: 10.1128/AEM.70.6.3493-3499.2004. Appl Environ Microbiol. 2004. PMID: 15184148 Free PMC article.
-
Molecular cloning and nucleotide sequence of the gene encoding the major peptidoglycan hydrolase of Lactococcus lactis, a muramidase needed for cell separation.J Bacteriol. 1995 Mar;177(6):1554-63. doi: 10.1128/jb.177.6.1554-1563.1995. J Bacteriol. 1995. PMID: 7883712 Free PMC article.
-
Characterization of AcmB, an N-acetylglucosaminidase autolysin from Lactococcus lactis.Microbiology (Reading). 2003 Mar;149(Pt 3):695-705. doi: 10.1099/mic.0.25875-0. Microbiology (Reading). 2003. PMID: 12634338
-
The autolytic ('suicidase') system of Enterococcus hirae: from lysine depletion autolysis to biochemical and molecular studies of the two muramidases of Enterococcus hirae ATCC 9790.FEMS Microbiol Lett. 1992 Dec 15;100(1-3):261-7. doi: 10.1111/j.1574-6968.1992.tb14050.x. FEMS Microbiol Lett. 1992. PMID: 1362171 Review.
-
The pneumococcal cell wall degrading enzymes: a modular design to create new lysins?Microb Drug Resist. 1997 Summer;3(2):199-211. doi: 10.1089/mdr.1997.3.199. Microb Drug Resist. 1997. PMID: 9185148 Review.
Cited by
-
Optimization of the Lactococcus lactis nisin-controlled gene expression system NICE for industrial applications.Microb Cell Fact. 2005 May 30;4:16. doi: 10.1186/1475-2859-4-16. Microb Cell Fact. 2005. PMID: 15921537 Free PMC article.
-
In vitro and in vivo probiotic assessment of Leuconostoc mesenteroides P45 isolated from pulque, a Mexican traditional alcoholic beverage.Springerplus. 2016 Jun 13;5(1):708. doi: 10.1186/s40064-016-2370-7. eCollection 2016. Springerplus. 2016. PMID: 27375977 Free PMC article.
-
Analysis of the peptidoglycan hydrolase complement of Lactococcus lactis: identification of a third N-acetylglucosaminidase, AcmC.Appl Environ Microbiol. 2004 Jun;70(6):3493-9. doi: 10.1128/AEM.70.6.3493-3499.2004. Appl Environ Microbiol. 2004. PMID: 15184148 Free PMC article.
-
Identification of the chain-dispersing peptidoglycan hydrolase LytB of Streptococcus gordonii.PLoS One. 2017 Apr 17;12(4):e0176117. doi: 10.1371/journal.pone.0176117. eCollection 2017. PLoS One. 2017. PMID: 28414782 Free PMC article.
-
Production of the Bacillus licheniformis SubC protease using Lactococcus lactis NICE expression system.Springerplus. 2012 Nov 29;1:54. doi: 10.1186/2193-1801-1-54. eCollection 2012. Springerplus. 2012. PMID: 23961373 Free PMC article.
References
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources

