Cloning of the sodA gene from Corynebacterium melassecola and role of superoxide dismutase in cellular viability
- PMID: 11157941
- PMCID: PMC95002
- DOI: 10.1128/JB.2001.183.4.1284-1295.2001
Cloning of the sodA gene from Corynebacterium melassecola and role of superoxide dismutase in cellular viability
Abstract
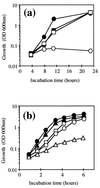
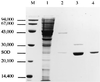
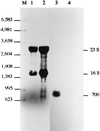
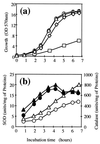
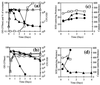
The sodA gene encoding the Corynebacterium melassecola manganese-cofactored superoxide dismutase (SOD) has been cloned in Escherichia coli and sequenced. The gene is transcribed monocistronically; the predicted polypeptide is 200 amino acids long and associates in a homotetrameric, manganese-dependent form, able to complement an SOD-deficient E. coli mutant. A second open reading frame, coding for a putative 217-amino-acid protein with high homology to peptide methionine sulfoxide reductases from various origins, has been identified immediately upstream of sodA in the opposite transcription orientation. The sodA gene was inactivated by insertion of an integrative vector carrying a kanamycin resistance gene. The growth rate of the SOD-deficient integrant was only slightly affected in BHI rich medium as well as in BMCG chemically defined medium, but was strongly affected by the presence of the redox-cycling agent paraquat. The SOD deficiency had, on the other hand, a deleterious effect on viability as soon as the culture entered the stationary phase of growth in BHI medium. Surprisingly, SOD deficiency was able to rescue the dramatic loss of viability observed for the wild-type strain in BMCG synthetic medium when glucose was not the limiting growth factor.
Figures

Similar articles
-
Cloning and characterization of the Pseudomonas aeruginosa sodA and sodB genes encoding manganese- and iron-cofactored superoxide dismutase: demonstration of increased manganese superoxide dismutase activity in alginate-producing bacteria.J Bacteriol. 1993 Dec;175(23):7658-65. doi: 10.1128/jb.175.23.7658-7665.1993. J Bacteriol. 1993. PMID: 8244935 Free PMC article.
-
Pseudomonas aeruginosa sodA and sodB mutants defective in manganese- and iron-cofactored superoxide dismutase activity demonstrate the importance of the iron-cofactored form in aerobic metabolism.J Bacteriol. 1995 Nov;177(22):6330-7. doi: 10.1128/jb.177.22.6330-6337.1995. J Bacteriol. 1995. PMID: 7592406 Free PMC article.
-
Characterization of the superoxide dismutase gene and its upstream region from Methanobacterium thermoautotrophicum Marburg.FEMS Microbiol Lett. 1995 May 15;128(3):247-53. doi: 10.1111/j.1574-6968.1995.tb07532.x. FEMS Microbiol Lett. 1995. PMID: 7781971
-
Cloning, nucleotide sequence and characterization of a gene encoding superoxide dismutase from Campylobacter jejuni and Campylobacter coli.Microbiology (Reading). 1994 May;140 ( Pt 5):1203-8. doi: 10.1099/13500872-140-5-1203. Microbiology (Reading). 1994. PMID: 8025686
-
Essential role of superoxide dismutase on the pathogenicity of Erwinia chrysanthemi strain 3937.Mol Plant Microbe Interact. 2001 Jun;14(6):758-67. doi: 10.1094/MPMI.2001.14.6.758. Mol Plant Microbe Interact. 2001. PMID: 11386371
Cited by
-
Corynebacterium diphtheriae Proteome Adaptation to Cell Culture Medium and Serum.Proteomes. 2021 Mar 13;9(1):14. doi: 10.3390/proteomes9010014. Proteomes. 2021. PMID: 33805816 Free PMC article.
-
Differential expression and localization of Mn and Fe superoxide dismutases in the heterocystous cyanobacterium Anabaena sp. strain PCC 7120.J Bacteriol. 2002 Sep;184(18):5096-103. doi: 10.1128/JB.184.18.5096-5103.2002. J Bacteriol. 2002. PMID: 12193626 Free PMC article.
-
The complete genome sequence of Corynebacterium pseudotuberculosis FRC41 isolated from a 12-year-old girl with necrotizing lymphadenitis reveals insights into gene-regulatory networks contributing to virulence.BMC Genomics. 2010 Dec 30;11:728. doi: 10.1186/1471-2164-11-728. BMC Genomics. 2010. PMID: 21192786 Free PMC article.
-
The superoxide dismutase gene sodM is unique to Staphylococcus aureus: absence of sodM in coagulase-negative staphylococci.J Bacteriol. 2002 May;184(9):2465-72. doi: 10.1128/JB.184.9.2465-2472.2002. J Bacteriol. 2002. PMID: 11948161 Free PMC article.
-
Inactivation of the Moraxella catarrhalis superoxide dismutase SodA induces constitutive expression of iron-repressible outer membrane proteins.Infect Immun. 2002 Apr;70(4):1889-95. doi: 10.1128/IAI.70.4.1889-1895.2002. Infect Immun. 2002. PMID: 11895952 Free PMC article.
References
-
- Alcendor D J, Chapman G D, Beaman B L. Isolation, sequencing and expression of the superoxide dismutase-encoding gene (sod) of Nocardia asteroides strain GUH-2. Gene. 1995;164:143–147. - PubMed
-
- Amador E, Castro J M, Correia A, Martin J F. Structure and organization of the rrnD operon of ‘Brevibacterium lactofermentum’: analysis of the 16S rRNA gene. Microbiology. 1999;145:915–924. - PubMed
-
- Battistoni A, Pacello F, Folcarelli S, Ajello M, Donnarumma G, Greco R, Ammendolia M G, Touati D, Rotilio G, Valenti P. Increased expression of periplasmic Cu,Zn superoxide dismutase enhances survival of Escherichia coli invasive strains within nonphagocytic cells. Infect Immun. 2000;68:30–37. - PMC - PubMed
-
- Beck B L, Tabatabai L B, Mayfield J E. A protein isolated from Brucella abortus is a Cu-Zn superoxide dismutase. Biochemistry. 1990;29:372–376. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources

