Forced unfolding modulated by disulfide bonds in the Ig domains of a cell adhesion molecule
- PMID: 11171991
- PMCID: PMC29297
- DOI: 10.1073/pnas.98.4.1565
Forced unfolding modulated by disulfide bonds in the Ig domains of a cell adhesion molecule
Abstract
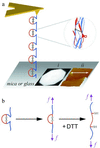
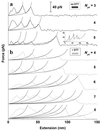
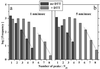
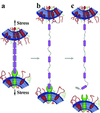
Cell adhesion molecules (CAMs) mediate cell attachment and stress transfer through extracellular domains. Here we forcibly unfold the Ig domains of a prototypical Ig superfamily CAM that contains intradomain disulfide bonds. The Ig domains of all such CAMs have conformations homologous to cadherin extracellular domains, titin Ig-type domains, and fibronectin type-III (FNIII) domains. Atomic force microscopy has been used to extend the five Ig domains of Mel-CAM (melanoma CAM)--a protein that is overexpressed in metastatic melanomas--under conditions where the disulfide bonds were either left intact or disrupted through reduction. Under physiological conditions where intradomain disulfide bonds are intact, partial unfolding was observed at forces far smaller than those reported previously for either titin's Ig-type domains or tenascin's FNIII domains. This partial unfolding under low force may be an important mechanism for imparting elasticity to cell-cell contacts, as well as a regulatory mechanism for adhesive interactions. Under reducing conditions, Mel-CAM's Ig domains were found to fully unfold through a partially folded state and at slightly higher forces. The results suggest that, in divergent evolution of all such domains, stabilization imparted by disulfide bonds relaxes requirements for strong, noncovalent, folded-state interactions.
Figures



References
-
- Thiery J P. In: Guidebook to Extracellular Matrix, Anchor, and Adhesion Proteins. Kreis T, Vale R, editors. New York: Oxford Univ. Press; 2000. pp. 125–128.
-
- Shih I M, Elder D E, Speicher D, Johnson J P, Herlyn M. Cancer Res. 1994;54:2514–2520. - PubMed
-
- Shih I M, Kurman R J. Lab Invest. 1996;75:377–388. - PubMed
-
- Pickl W F, Majdic O, Fischer G F, Petzelbauer P, Fae I, Waclavicek M, Stockl J, Scheinecker C, Vidicki T, Aschauer H, et al. J Immunol. 1997;158:2107–2115. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

