Two-component sensor required for normal symbiotic colonization of euprymna scolopes by Vibrio fischeri
- PMID: 11208780
- PMCID: PMC94949
- DOI: 10.1128/JB.183.3.835-842.2001
Two-component sensor required for normal symbiotic colonization of euprymna scolopes by Vibrio fischeri
Abstract
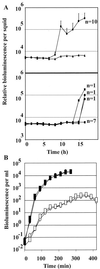
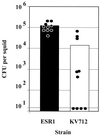
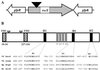
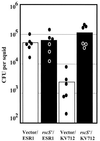
The light organ of the squid Euprymna scolopes is specifically colonized to a high density by the marine bacterium Vibrio fischeri. To date, only a few factors contributing to the specificity of this symbiosis have been identified. Using a genetic screen for random transposon mutants defective in initiating the symbiotic association or in colonizing the light organ to high density, we identified a mutant of V. fischeri that exhibited an apparent defect in symbiosis initiation. This mutant was not defective in motility, luminescence, or growth in minimal medium, suggesting that it lacks an essential, previously unidentified symbiotic function. By sequence analysis, we showed that the locus inactivated in this mutant encodes a predicted 927-amino-acid protein with a high degree of similarity to the sensor component of hybrid two-component regulatory systems. We have therefore designated this locus rscS, for regulator of symbiotic colonization-sensor. Sequence analysis revealed two hydrophobic regions which may result in the formation of a periplasmic loop involved in signal recognition; PhoA fusion data supported this proposed membrane topology. We have investigated the start site of rscS transcription by primer extension and identified a putative promoter region. We hypothesize that RscS recognizes a signal associated with the light organ environment and responds by stimulating a putative response regulator that controls protein function or gene expression to coordinate early colonization events. Further studies on RscS, its cognate response regulator, and the signaling conditions will provide important insight into the interaction between V. fischeri and E. scolopes.
Figures

References
-
- Alexeyev M F, Shokolenko I N. Mini-Tn10 transposon derivatives for insertion mutagenesis and gene delivery into the chromosome of Gram-negative bacteria. Gene. 1995;160:59–62. - PubMed
-
- Appleby J L, Parkinson J S, Bourret R B. Signal transduction via the multi-step phosphorelay: not necessarily a road less traveled. Cell. 1996;86:845–848. - PubMed
-
- Arico B, Scarlato V, Monack D M, Falkow S, Rappuoli R. Structural and genetic analysis of the bvg locus in Bordetella species. Mol Microbiol. 1991;5:2481–2491. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources

