Characterization and mutational analysis of yeast Dbp8p, a putative RNA helicase involved in ribosome biogenesis
- PMID: 11222764
- PMCID: PMC29721
- DOI: 10.1093/nar/29.5.1144
Characterization and mutational analysis of yeast Dbp8p, a putative RNA helicase involved in ribosome biogenesis
Abstract
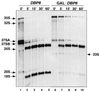
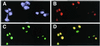
RNA helicases of the DEAD box family are involved in almost all cellular processes involving RNA molecules. Here we describe functional characterization of the yeast RNA helicase Dbp8p (YHR169w). Our results show that Dbp8p is an essential nucleolar protein required for biogenesis of the small ribosomal subunit. In vivo depletion of Dbp8p resulted in a ribosomal subunit imbalance due to a deficit in 40S ribosomal subunits. Subsequent analyses of pre-rRNA processing by pulse-chase labeling, northern hybridization and primer extension revealed that the early steps of cleavage of the 35S precursor at sites A(1) and A(2) are inhibited and delayed at site A(0). Synthesis of 18S rRNA, the RNA moiety of the 40S subunit, is thereby blocked in the absence of Dbp8p. The involvement of Dbp8p as a bona fide RNA helicase in ribosome biogenesis is strongly supported by the loss of Dbp8p in vivo function obtained by site-directed mutagenesis of some conserved motifs carrying the enzymatic properties of the protein family.
Figures







Similar articles
-
Has1p, a member of the DEAD-box family, is required for 40S ribosomal subunit biogenesis in Saccharomyces cerevisiae.Mol Microbiol. 2004 Apr;52(1):141-58. doi: 10.1111/j.1365-2958.2003.03973.x. Mol Microbiol. 2004. PMID: 15049817
-
Fal1p is an essential DEAD-box protein involved in 40S-ribosomal-subunit biogenesis in Saccharomyces cerevisiae.Mol Cell Biol. 1997 Dec;17(12):7283-94. doi: 10.1128/MCB.17.12.7283. Mol Cell Biol. 1997. PMID: 9372960 Free PMC article.
-
Dbp3p, a putative RNA helicase in Saccharomyces cerevisiae, is required for efficient pre-rRNA processing predominantly at site A3.Mol Cell Biol. 1997 Mar;17(3):1354-65. doi: 10.1128/MCB.17.3.1354. Mol Cell Biol. 1997. PMID: 9032262 Free PMC article.
-
Unwinding RNA in Saccharomyces cerevisiae: DEAD-box proteins and related families.Trends Biochem Sci. 1999 May;24(5):192-8. doi: 10.1016/s0968-0004(99)01376-6. Trends Biochem Sci. 1999. PMID: 10322435 Review.
-
DExD/H-box RNA helicases in ribosome biogenesis.RNA Biol. 2013 Jan;10(1):4-18. doi: 10.4161/rna.21879. Epub 2012 Aug 24. RNA Biol. 2013. PMID: 22922795 Free PMC article. Review.
Cited by
-
RNA folding and functions of RNA helicases in ribosome biogenesis.RNA Biol. 2022 Jan;19(1):781-810. doi: 10.1080/15476286.2022.2079890. RNA Biol. 2022. PMID: 35678541 Free PMC article. Review.
-
Characterization of the ATPase and unwinding activities of the yeast DEAD-box protein Has1p and the analysis of the roles of the conserved motifs.Nucleic Acids Res. 2005 Feb 17;33(3):999-1009. doi: 10.1093/nar/gki244. Print 2005. Nucleic Acids Res. 2005. PMID: 15718299 Free PMC article.
-
The nucleolar protein Esf2 interacts directly with the DExD/H box RNA helicase, Dbp8, to stimulate ATP hydrolysis.Nucleic Acids Res. 2006 Jun 13;34(10):3189-99. doi: 10.1093/nar/gkl419. Print 2006. Nucleic Acids Res. 2006. PMID: 16772403 Free PMC article.
-
Differential RNA-dependent ATPase activities of four rRNA processing yeast DEAD-box proteins.Biochemistry. 2008 Nov 25;47(47):12562-73. doi: 10.1021/bi8016119. Biochemistry. 2008. PMID: 18975973 Free PMC article.
-
Unraveling the 'DEAD-box' helicases of Plasmodium falciparum.Gene. 2006 Jul 5;376(1):1-12. doi: 10.1016/j.gene.2006.03.007. Epub 2006 Apr 7. Gene. 2006. PMID: 16713133 Free PMC article. Review.
References
-
- Linder P., Lasko,P.F., Ashburner,M., Leroy,P., Nielsen,P.J., Nishi,K., Schnier,J. and Slonimski,P.P. (1989) Birth of the D-E-A-D box. Nature, 337, 121–122. - PubMed
-
- de la Cruz J., Kressler,D. and Linder,P. (1999) Unwinding RNA in Saccharomyces cerevisiae: DEAD-box proteins and related families. Trends Biochem. Sci., 24, 192–198. - PubMed
-
- Iost I., Dreyfus,M. and Linder,P. (1999) Ded1p, a DEAD-box protein required for translation initation in Saccharomyces cerevisiae, is an RNA helicase. J. Biol. Chem., 274, 17677–17683. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

