Annealing of RNA editing substrates facilitated by guide RNA-binding protein gBP21
- PMID: 11250905
- PMCID: PMC145538
- DOI: 10.1093/emboj/20.6.1394
Annealing of RNA editing substrates facilitated by guide RNA-binding protein gBP21
Abstract
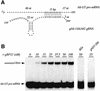
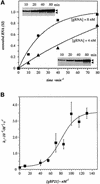
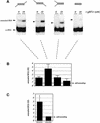
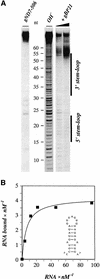
RNA editing within the mitochondria of African trypanosomes is characterized by the insertion and deletion of uridylate residues into otherwise incomplete primary transcripts. The reaction takes place in a high molecular mass ribonucleoprotein (RNP) complex of uncertain composition. Furthermore, factors that interact with the RNP complex during the reaction are by and large unknown. Here we present evidence for an editing-related biochemical activity of the gRNA-binding protein gBP21. Using recombinant gBP21 preparations, we show that the protein stimulates the annealing of gRNAs to cognate pre-mRNAs in vitro. This represents the presumed first step of the editing reaction. Kinetic data establish an enhancement of the second order rate constant for the gRNA- pre-mRNA interaction. gBP21-mediated annealing is not exclusive for RNA editing substrates since complementary RNAs, unrelated to the editing process, can also be hybridized. The gBP21-dependent RNA annealing activity was identified in mitochondrial extracts of trypanosomes and can be inhibited by immunoprecipitation of the polypeptide. The data suggest a factor-like contribution of gBP21 to the RNA editing process by accelerating the rate of gRNA-pre-mRNA anchor formation.
Figures



Similar articles
-
Trypanosoma brucei gBP21. An arginine-rich mitochondrial protein that binds to guide RNA with high affinity.J Biol Chem. 1997 Feb 7;272(6):3749-57. doi: 10.1074/jbc.272.6.3749. J Biol Chem. 1997. PMID: 9013632
-
The involvement of gRNA-binding protein gBP21 in RNA editing-an in vitro and in vivo analysis.Nucleic Acids Res. 1999 Mar 15;27(6):1429-36. doi: 10.1093/nar/27.6.1429. Nucleic Acids Res. 1999. PMID: 10037802 Free PMC article.
-
Cloning and characterization of two guide RNA-binding proteins from mitochondria of Crithidia fasciculata: gBP27, a novel protein, and gBP29, the orthologue of Trypanosoma brucei gBP21.Nucleic Acids Res. 2001 Jul 15;29(14):2950-62. doi: 10.1093/nar/29.14.2950. Nucleic Acids Res. 2001. PMID: 11452020 Free PMC article.
-
Insertional and deletional RNA editing in trypanosome mitochondria.Nucleic Acids Symp Ser. 1997;(36):15-8. Nucleic Acids Symp Ser. 1997. PMID: 9478193 Review.
-
Mitochondrial RNA editing in trypanosomes: small RNAs in control.Biochimie. 2014 May;100:125-31. doi: 10.1016/j.biochi.2014.01.003. Epub 2014 Jan 17. Biochimie. 2014. PMID: 24440637 Free PMC article. Review.
Cited by
-
Intrinsic and regulated properties of minimally edited trypanosome mRNAs.Nucleic Acids Res. 2019 Apr 23;47(7):3640-3657. doi: 10.1093/nar/gkz012. Nucleic Acids Res. 2019. PMID: 30698753 Free PMC article.
-
Transient RNA-protein interactions in RNA folding.FEBS J. 2011 May;278(10):1634-42. doi: 10.1111/j.1742-4658.2011.08094.x. Epub 2011 Apr 13. FEBS J. 2011. PMID: 21410645 Free PMC article. Review.
-
Minimal pre-mRNA substrates with natural and converted sites for full-round U insertion and U deletion RNA editing in trypanosomes.Nucleic Acids Res. 2005 Nov 23;33(20):6610-20. doi: 10.1093/nar/gki943. Print 2005. Nucleic Acids Res. 2005. PMID: 16306234 Free PMC article.
-
Compositionally and functionally distinct editosomes in Trypanosoma brucei.RNA. 2006 Jun;12(6):1038-49. doi: 10.1261/rna.45506. Epub 2006 Apr 12. RNA. 2006. PMID: 16611942 Free PMC article.
-
Pentatricopeptide repeat poly(A) binding protein KPAF4 stabilizes mitochondrial mRNAs in Trypanosoma brucei.Nat Commun. 2019 Jan 11;10(1):146. doi: 10.1038/s41467-018-08137-2. Nat Commun. 2019. PMID: 30635574 Free PMC article.
References
-
- Benne R. and Speijer,D. (1998) RNA editing types and characteristics. In Grosjean,H. and Benne,R. (eds), Modification and Editing of RNA. ASM Press, Washington, DC, pp. 551–554.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

