The human XPG gene: gene architecture, alternative splicing and single nucleotide polymorphisms
- PMID: 11266544
- PMCID: PMC31292
- DOI: 10.1093/nar/29.7.1443
The human XPG gene: gene architecture, alternative splicing and single nucleotide polymorphisms
Abstract
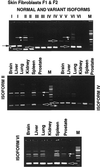
Defects in the XPG DNA repair endonuclease gene can result in the cancer-prone disorders xeroderma pigmentosum (XP) or the XP-Cockayne syndrome complex. While the XPG cDNA sequence was known, determination of the genomic sequence was required to understand its different functions. In cells from normal donors, we found that the genomic sequence of the human XPG gene spans 30 kb, contains 15 exons that range from 61 to 1074 bp and 14 introns that range from 250 to 5763 bp. Analysis of the splice donor and acceptor sites using an information theory-based approach revealed three splice sites with low information content, which are components of the minor (U12) spliceosome. We identified six alternatively spliced XPG mRNA isoforms in cells from normal donors and from XPG patients: partial deletion of exon 8, partial retention of intron 8, two with alternative exons (in introns 1 and 6) and two that retained complete introns (introns 3 and 9). The amount of alternatively spliced XPG mRNA isoforms varied in different tissues. Most alternative splice donor and acceptor sites had a relatively high information content, but one has the U12 spliceosome sequence. A single nucleotide polymorphism has allele frequencies of 0.74 for 3507G and 0.26 for 3507C in 91 donors. The human XPG gene contains multiple splice sites with low information content in association with multiple alternatively spliced isoforms of XPG mRNA.
Figures


References
-
- Bootsma D., Kraemer,K.H., Cleaver,J.E. and Hoeijmakers,J.H. (1998) Nucleotide excision repair syndromes: xeroderma pigmentosum, Cockayne syndrome, and trichothiodystrophy. In Vogelstein,B. and Kinzler,K.W. (eds), The Genetic Basis of Human Cancer. McGraw-Hill, New York, NY, pp. 245–274.
-
- Kraemer K.H., Lee,M.M. and Scotto,J. (1987) Xeroderma pigmentosum. Cutaneous, ocular, and neurologic abnormalities in 830 published cases. Arch. Dermatol., 123, 241–250. - PubMed
-
- Kraemer K.H., Lee,M.M., Andrews,A.D. and Lambert,W.C. (1994) The role of sunlight and DNA repair in melanoma and nonmelanoma skin cancer. The xeroderma pigmentosum paradigm. Arch. Dermatol., 130, 1018–1021. - PubMed
-
- van Steeg H. and Kraemer,K.H. (1999) Xeroderma pigmentosum and the role of UV-induced DNA damage in skin cancer. Mol. Med. Today, 5, 86–94. - PubMed
-
- Masutani C., Kusumoto,R., Yamada,A., Dohmae,N., Yokoi,M., Yuasa,M., Araki,M., Iwai,S., Takio,K. and Hanaoka,F. (1999) The XPV (xeroderma pigmentosum variant) gene encodes human DNA polymerase η. Nature, 399, 700–704. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

