Regulation of DAF-2 receptor signaling by human insulin and ins-1, a member of the unusually large and diverse C. elegans insulin gene family
- PMID: 11274053
- PMCID: PMC312654
- DOI: 10.1101/gad.867301
Regulation of DAF-2 receptor signaling by human insulin and ins-1, a member of the unusually large and diverse C. elegans insulin gene family
Abstract
The activity of the DAF-2 insulin-like receptor is required for Caenorhabditis elegans reproductive growth and normal adult life span. Informatic analysis identified 37 C. elegans genes predicted to encode insulin-like peptides. Many of these genes are divergent insulin superfamily members, and many are clustered, indicating recent diversification of the family. The ins genes are primarily expressed in neurons, including sensory neurons, a subset of which are required for reproductive development. Structural predictions and likely C-peptide cleavage sites typical of mammalian insulins suggest that ins-1 is most closely related to insulin. Overexpression of ins-1, or expression of human insulin under the control of ins-1 regulatory sequences, causes partially penetrant arrest at the dauer stage and enhances dauer arrest in weak daf-2 mutants, suggesting that INS-1 and human insulin antagonize DAF-2 insulin-like signaling. A deletion of the ins-1 coding region does not enhance or suppress dauer arrest, indicating a functional redundancy among the 37 ins genes. Of five other ins genes tested, the only other one bearing a predicted C peptide also antagonizes daf-2 signaling, whereas four ins genes without a C peptide do not, indicating functional diversity within the ins family.
Figures


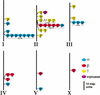
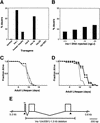

References
-
- Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215:403–410. - PubMed
-
- Baker EN, Blundell TL, Cutfield JF, Cutfield SM, Dodson EJ, Dodson GG, Hodgkin DM, Hubbard RE, Isaacs NW, Reynolds CD, et al. The structure of 2Zn pig insulin crystals at 1.5 Å resolution. Philos Trans R Soc Lond B Biol Sci. 1988;319:369–456. - PubMed
-
- Bargmann CI, Horvitz HR. Control of larval development by chemosensory neurons in Caenorhabditis elegans. Science. 1991;251:1243–1246. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous
