Free energy reconstruction from nonequilibrium single-molecule pulling experiments
- PMID: 11274384
- PMCID: PMC31107
- DOI: 10.1073/pnas.071034098
Free energy reconstruction from nonequilibrium single-molecule pulling experiments
Abstract
Laser tweezers and atomic force microscopes are increasingly used to probe the interactions and mechanical properties of individual molecules. Unfortunately, using such time-dependent perturbations to force rare molecular events also drives the system away from equilibrium. Nevertheless, we show how equilibrium free energy profiles can be extracted rigorously from repeated nonequilibrium force measurements on the basis of an extension of Jarzynski's remarkable identity between free energies and the irreversible work.
Figures

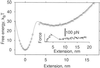

Comment in
-
How does a system respond when driven away from thermal equilibrium?Proc Natl Acad Sci U S A. 2001 Mar 27;98(7):3636-8. doi: 10.1073/pnas.081074598. Proc Natl Acad Sci U S A. 2001. PMID: 11274379 Free PMC article. No abstract available.
Similar articles
-
Free energy surfaces from single-molecule force spectroscopy.Acc Chem Res. 2005 Jul;38(7):504-13. doi: 10.1021/ar040148d. Acc Chem Res. 2005. PMID: 16028884
-
Deciphering the scaling of single-molecule interactions using Jarzynski's equality.Nat Commun. 2014 Nov 21;5:5539. doi: 10.1038/ncomms6539. Nat Commun. 2014. PMID: 25412574
-
Hummer and Szabo-like potential of mean force estimator for bidirectional nonequilibrium pulling experiments/simulations.J Phys Chem B. 2010 Jul 29;114(29):9546-54. doi: 10.1021/jp102263y. J Phys Chem B. 2010. PMID: 20597536
-
Velocity convergence of free energy surfaces from single-molecule measurements using Jarzynski's equality.Phys Rev E Stat Nonlin Soft Matter Phys. 2009 Apr;79(4 Pt 1):041912. doi: 10.1103/PhysRevE.79.041912. Epub 2009 Apr 10. Phys Rev E Stat Nonlin Soft Matter Phys. 2009. PMID: 19518261
-
Extracting physics of life at the molecular level: A review of single-molecule data analyses.Phys Life Rev. 2015 Jun;13:107-37. doi: 10.1016/j.plrev.2015.01.017. Epub 2015 Jan 14. Phys Life Rev. 2015. PMID: 25660417 Review.
Cited by
-
Mechanism of titin unfolding by force: insight from quasi-equilibrium molecular dynamics calculations.Biophys J. 2006 Jul 15;91(2):467-72. doi: 10.1529/biophysj.106.082594. Epub 2006 Apr 21. Biophys J. 2006. PMID: 16632514 Free PMC article.
-
A Companion Guide to the String Method with Swarms of Trajectories: Characterization, Performance, and Pitfalls.J Chem Theory Comput. 2022 Mar 8;18(3):1406-1422. doi: 10.1021/acs.jctc.1c01049. Epub 2022 Feb 9. J Chem Theory Comput. 2022. PMID: 35138832 Free PMC article.
-
Free energy of membrane protein unfolding derived from single-molecule force measurements.Biophys J. 2007 Aug 1;93(3):930-7. doi: 10.1529/biophysj.106.096982. Epub 2007 May 4. Biophys J. 2007. PMID: 17483176 Free PMC article.
-
Renormalizing SMD: the renormalization approach and its use in long time simulations and accelerated PMF calculations of macromolecules.J Phys Chem B. 2010 Oct 7;114(39):12720-8. doi: 10.1021/jp1056122. J Phys Chem B. 2010. PMID: 20836533 Free PMC article.
-
Antibody-nanobody combination increases their neutralizing activity against SARS-CoV-2 and nanobody H11-H4 is effective against Alpha, Kappa and Delta variants.Sci Rep. 2022 Jun 11;12(1):9701. doi: 10.1038/s41598-022-14263-1. Sci Rep. 2022. PMID: 35690632 Free PMC article.
References
-
- Perkins T T, Smith D E, Chu S. Science. 1994;264:819–822. - PubMed
-
- Florin E L, Moy V T, Gaub H E. Science. 1994;264:415–417. - PubMed
-
- Strick T R, Allemand J F, Bensimon D, Bensimon A, Croquette V. Science. 1996;271:1835–1837. - PubMed
-
- Smith S B, Cui Y J, Bustamante C. Science. 1996;271:795–799. - PubMed
-
- Tskhovrebova L, Trinick J, Sleep J A, Simmons R M. Nature (London) 1997;387:308–312. - PubMed
LinkOut - more resources
Full Text Sources

