Visualization of oligonucleotide probes and point mutations in interphase nuclei and DNA fibers using rolling circle DNA amplification
- PMID: 11274414
- PMCID: PMC31158
- DOI: 10.1073/pnas.061026198
Visualization of oligonucleotide probes and point mutations in interphase nuclei and DNA fibers using rolling circle DNA amplification
Abstract
Rolling circle amplification (RCA) is a surface-anchored DNA replication reaction that can be exploited to visualize single molecular recognition events. Here we report the use of RCA to visualize target DNA sequences as small as 50 nts in peripheral blood lymphocytes or in stretched DNA fibers. Three unique target sequences within the cystic fibrosis transmembrane conductance regulator gene could be detected simultaneously in interphase nuclei, and could be ordered in a linear map in stretched DNA. Allele-discriminating oligonucleotide probes in conjunction with RCA also were used to discriminate wild-type and mutant alleles in the cystic fibrosis transmembrane conductance regulator, p53, BRCA-1, and Gorlin syndrome genes in the nuclei of cultured cells or in DNA fibers. These observations demonstrate that signal amplification by RCA can be coupled to nucleic acid hybridization and multicolor fluorescence imaging to detect single nucleotide changes in DNA within a cytological context or in single DNA molecules. This provides a means for direct physical haplotyping and the analysis of somatic mutations on a cell-by-cell basis.
Figures
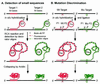



References
-
- Nath J, Johnson K L. Biotechnol Histochem. 2000;75:54–78. - PubMed
-
- Raap A K. Mutat Res. 1998;400:287–298. - PubMed
-
- Van Gijlswijk R P, van de Corput M P, Begrookove V, Wiegart J, Tanke H J, Raap A. Histochem Cell Biol. 2000;113:175–180. - PubMed
-
- Lizardi P M, Huang X, Zhu Z, Bray-Ward P, Thomas D C, Ward D C. Nat Genet. 1998;19:225–232. - PubMed
-
- Thomas D C, Nardone G A, Randall S K. Arch Pathol Lab Med. 1999;123:1170–1176. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

