Marek's disease virus (MDV) encodes an interleukin-8 homolog (vIL-8): characterization of the vIL-8 protein and a vIL-8 deletion mutant MDV
- PMID: 11333897
- PMCID: PMC114921
- DOI: 10.1128/JVI.75.11.5159-5173.2001
Marek's disease virus (MDV) encodes an interleukin-8 homolog (vIL-8): characterization of the vIL-8 protein and a vIL-8 deletion mutant MDV
Abstract
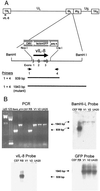
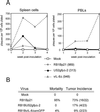
Chemokines induce chemotaxis, cell migration, and inflammatory responses. We report the identification of an interleukin-8 (IL-8) homolog, termed vIL-8, encoded within the genome of Marek's disease virus (MDV). The 134-amino-acid vIL-8 shares closest homology to mammalian and avian IL-8, molecules representing the prototype CXC chemokine. The gene for vIL-8 consists of three exons which map to the BamHI-L fragment within the repeats flanking the unique long region of the MDV genome. A 0.7-kb transcript encoding vIL-8 was detected in an n-butyrate-treated, MDV-transformed T-lymphoblastoid cell line, MSB-1. This induction is essentially abolished by cycloheximide and herpesvirus DNA polymerase inhibitor phosphonoacetate, indicating that vIL-8 is expressed with true late (gamma2) kinetics. Baculovirus-expressed vIL-8 was found to be secreted into the medium and shown to be functional as a chemoattractant for chicken peripheral blood mononuclear cells but not for heterophils. To characterize the function of vIL-8 with respect to MDV infection in vivo, a recombinant MDV was constructed with a deletion of all three exons and a soluble-modified green fluorescent protein (smGFP) expression cassette inserted at the site of deletion. In two in vivo experiments, the vIL-8 deletion mutant (RB1BvIL-8DeltasmGFP) showed a decreased level of lytic infection in comparison to its parent virus, an equal-passage-level parent virus, and to another recombinant MDV containing the insertion of a GFP expression cassette at the nonessential US2 gene. RB1BvIL-8DeltasmGFP retained oncogenicity, albeit at a greatly reduced level. Nonetheless, we have been able to establish a lymphoblastoid cell line from an RB1BvIL-8DeltasmGFP-induced ovarian lymphoma (MDCC-UA20) and verify the presence of a latent MDV genome lacking vIL-8. Taken together, these data describe the identification and characterization of a chemokine homolog encoded within the MDV genome that is dispensable for transformation but may affect the level of MDV in vivo lytic infection.
Figures





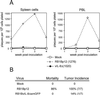

Similar articles
-
Impact of deletions within the Bam HI-L fragment of attenuated Marek's disease virus on vIL-8 expression and the newly identified transcript of open reading frame LORF4.Virus Genes. 2003 May;26(3):255-69. doi: 10.1023/a:1024447230464. Virus Genes. 2003. PMID: 12876454
-
Marek's disease virus-encoded vIL-8 gene is involved in early cytolytic infection but dispensable for establishment of latency.J Virol. 2004 May;78(9):4753-60. doi: 10.1128/jvi.78.9.4753-4760.2004. J Virol. 2004. PMID: 15078957 Free PMC article.
-
Construction and characterization of Marek's disease viruses having green fluorescent protein expression tied directly or indirectly to phosphoprotein 38 expression.Avian Dis. 2004 Sep;48(3):471-87. doi: 10.1637/7110. Avian Dis. 2004. PMID: 15529969
-
Latency and tumorigenesis in Marek's disease.Avian Dis. 2013 Jun;57(2 Suppl):360-5. doi: 10.1637/10470-121712-Reg.1. Avian Dis. 2013. PMID: 23901747 Review.
-
Marek's disease virus reactivation from latency: changes in gene expression at the origin of replication.Poult Sci. 2003 Jun;82(6):893-8. doi: 10.1093/ps/82.6.893. Poult Sci. 2003. PMID: 12817443 Review.
Cited by
-
Marek's disease viral interleukin-8 promotes lymphoma formation through targeted recruitment of B cells and CD4+ CD25+ T cells.J Virol. 2012 Aug;86(16):8536-45. doi: 10.1128/JVI.00556-12. Epub 2012 May 30. J Virol. 2012. PMID: 22647701 Free PMC article.
-
Fully Attenuated meq and pp38 Double Gene Deletion Mutant Virus Confers Superior Immunological Protection against Highly Virulent Marek's Disease Virus Infection.Microbiol Spectr. 2022 Dec 21;10(6):e0287122. doi: 10.1128/spectrum.02871-22. Epub 2022 Nov 9. Microbiol Spectr. 2022. PMID: 36350141 Free PMC article.
-
Histone modifications induced by MDV infection at early cytolytic and latency phases.BMC Genomics. 2015 Apr 18;16(1):311. doi: 10.1186/s12864-015-1492-6. BMC Genomics. 2015. PMID: 25896894 Free PMC article.
-
Impact of deletions within the Bam HI-L fragment of attenuated Marek's disease virus on vIL-8 expression and the newly identified transcript of open reading frame LORF4.Virus Genes. 2003 May;26(3):255-69. doi: 10.1023/a:1024447230464. Virus Genes. 2003. PMID: 12876454
-
Imaging Mass Spectrometry and Proteome Analysis of Marek's Disease Virus-Induced Tumors.mSphere. 2019 Jan 16;4(1):e00569-18. doi: 10.1128/mSphere.00569-18. mSphere. 2019. PMID: 30651403 Free PMC article.
References
-
- Adams D H, Lloyd A R. Chemokines: leucocyte recruitment and activation cytokines. Lancet. 1997;349:490–495. - PubMed
-
- Adams D H, Shaw S. Leucocyte-endothelial interactions and regulation of leucocyte migration. Lancet. 1994;343:831–836. - PubMed
-
- Akiyama Y, Kato S, Iwa N. Continuous cell culture from lymphoma of Marek's disease. Biken J. 1973;16:177–179. - PubMed
-
- Ausubel F M, Brent R, Kingston R E, Moore D D, Seidman J G, Smith J A, Struhl K. Current protocols in molecular biology. New York, N.Y: John Wiley & Sons; 1995.
-
- Baggiolini M. Chemokines and leukocyte traffic. Nature. 1998;392:565–568. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

