Incoming nucleotide binds to Klenow ternary complex leading to stable physical sequestration of preceding dNTP on DNA
- PMID: 11353079
- PMCID: PMC55446
- DOI: 10.1093/nar/29.10.2097
Incoming nucleotide binds to Klenow ternary complex leading to stable physical sequestration of preceding dNTP on DNA
Abstract
Klenow-DNA complex is known to undergo a rate-limiting, protein conformational transition from an 'open' to 'closed' state, upon binding of the 'correct' dNTP at the active site. In the 'closed' state, Mg(2+) mediates a rapid chemical step involving nucleophilic displacement of pyrophosphate by the 3' hydroxyl of the primer terminus. The enzyme returns to the 'open' state upon the release of PPi and translocation permits the next round of reaction. To determine whether Klenow can translocate to the next site on the addition of the next dNTP, without the preceding chemical step, we studied the ternary complex (Klenow-DNA-dNTP) in the absence of Mg(2+). While the ternary complex is proficient in chemical addition of dNTPs in Mg(2+), as revealed by primer extensions, the same in Mg(2+)-deficient conditions lead to non-covalent (physical) sequestration of first two 'correct' dNTPs in the ternary complex. Moreover, the second dNTP traps the first one in the DNA-helix of the ternary complex. Such a dNTP-DNA complex is found to be stable even after the dissociation of KLENOW: This reveals the novel state of the dNTP-DNA complex where the complementary base is stacked in a DNA-helix non-covalently, without the phosphodiester linkage. Further, shuttling of the DNA between the polymerase and the exonuclease site mediates the release of such a DNA complex. Interestingly, Klenow in such a Mg(2+)-deficient ternary complex exhibits a 'closed' conformation.
Figures



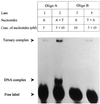





References
-
- Bryant F.R., Johnson,K.A. and Benkovic,S.J. (1983) Elementary steps in the DNA polymerase I reaction pathway. Biochemistry, 22, 3537–3546. - PubMed
-
- Mizrahi V., Henrie,R.N., Marlier,J.F., Johnson,K.A. and Benkovic,S.J. (1985) Rate limiting steps in the DNA polymerase I reaction pathway. Biochemistry, 24, 4010–4018. - PubMed
-
- Kuchta R.D., Mizrahi,V., Benkovic,P.A., Johnson,K.A. and Benkovic,S.J. (1987) Kinetic mechanism of DNA polymerase I (Klenow). Biochemistry, 26, 8410–8417. - PubMed
-
- Kuchta R.D., Benkovic,P.A. and Benkovic,S.J. (1988) Kinetic mechanism whereby DNA polymerase I (Klenow) replicates DNA with high fidelity. Biochemistry, 27, 6716–6725. - PubMed
-
- Eger B.T., Kuchta,R.D., Carroll,S.S., Benkovic,P.A., Dahlberg,M.E., Joyce,C.M. and Benkovic,S.J. (1991) Mechanism of DNA replication fidelity for the three mutants of DNA polymerase I: Klenow fragment KF (exo+), KF (polA5), and KF (exo–). Biochemistry, 30, 1441–1448. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

