Role of nuclear pools of aminoacyl-tRNA synthetases in tRNA nuclear export
- PMID: 11359929
- PMCID: PMC34591
- DOI: 10.1091/mbc.12.5.1381
Role of nuclear pools of aminoacyl-tRNA synthetases in tRNA nuclear export
Abstract
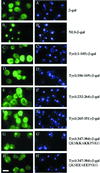
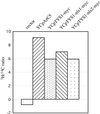
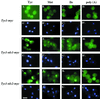
Reports of nuclear tRNA aminoacylation and its role in tRNA nuclear export (Lund and Dahlberg, 1998; Sarkar et al., 1999; Grosshans et al., 20001) have led to the prediction that there should be nuclear pools of aminoacyl-tRNA synthetases. We report that in budding yeast there are nuclear pools of tyrosyl-tRNA synthetase, Tys1p. By sequence alignments we predicted a Tys1p nuclear localization sequence and showed it to be sufficient for nuclear location of a passenger protein. Mutations of this nuclear localization sequence in endogenous Tys1p reduce nuclear Tys1p pools, indicating that the motif is also important for nucleus location. The mutations do not significantly affect catalytic activity, but they do cause defects in export of tRNAs to the cytosol. Despite export defects, the cells are viable, indicating that nuclear tRNA aminoacylation is not required for all tRNA nuclear export paths. Because the tRNA nuclear exportin, Los1p, is also unessential, we tested whether tRNA aminoacylation and Los1p operate in alternative tRNA nuclear export paths. No genetic interactions between aminoacyl-tRNA synthetases and Los1p were detected, indicating that tRNA nuclear aminoacylation and Los1p operate in the same export pathway or there are more than two pathways for tRNA nuclear export.
Figures





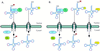
References
-
- Amberg DC, Goldstein AL, Cole CN. Isolation and characterization of RAT1: an essential gene of Saccharomyces cerevisiae required for the efficient nucleocytoplasmic trafficking of mRNA. Genes Dev. 1992;6:1173–1189. - PubMed
-
- Arts GJ, Fornerod M, Mattaj IW. Identification of a nuclear export receptor for tRNA. Curr Biol. 1998a;8:305–314. - PubMed
-
- Barbarese E, Koppel DE, Deutscher MP, Smith CL, Ainger K, Morgan F, Carson JH. Protein translation components are colocalized in granules in oligodendrocytes. J Cell Sci. 1995;108:2781–2790. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

