A dual role for substrate S-adenosyl-L-methionine in the methylation reaction with bacteriophage T4 Dam DNA-[N6-adenine]-methyltransferase
- PMID: 11376154
- PMCID: PMC55703
- DOI: 10.1093/nar/29.11.2361
A dual role for substrate S-adenosyl-L-methionine in the methylation reaction with bacteriophage T4 Dam DNA-[N6-adenine]-methyltransferase
Abstract
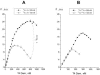
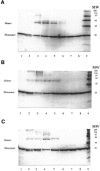
The fluorescence of 2-aminopurine ((2)A)-substituted duplexes (contained in the GATC target site) was investigated by titration with T4 Dam DNA-(N6-adenine)-methyltransferase. With an unmethylated target ((2)A/A duplex) or its methylated derivative ((2)A/(m)A duplex), T4 Dam produced up to a 50-fold increase in fluorescence, consistent with (2)A being flipped out of the DNA helix. Though neither S-adenosyl-L-homocysteine nor sinefungin had any significant effect, addition of substrate S-adenosyl-L-methionine (AdoMet) sharply reduced the Dam-induced fluorescence with these complexes. In contrast, AdoMet had no effect on the fluorescence increase produced with an (2)A/(2)A double-substituted duplex. Since the (2)A/(m)A duplex cannot be methylated, the AdoMet-induced decrease in fluorescence cannot be due to methylation per se. We propose that T4 Dam alone randomly binds to the asymmetric (2)A/A and (2)A/(m)A duplexes, and that AdoMet induces an allosteric T4 Dam conformational change that promotes reorientation of the enzyme to the strand containing the native base. Thus, AdoMet increases enzyme binding-specificity, in addition to serving as the methyl donor. The results of pre-steady-state methylation kinetics are consistent with this model.
Figures
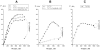
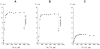
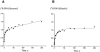
Similar articles
-
Bacteriophage T4 Dam DNA-[N6-adenine]methyltransferase. Kinetic evidence for a catalytically essential conformational change in the ternary complex.J Biol Chem. 2002 Jan 4;277(1):279-86. doi: 10.1074/jbc.M108864200. Epub 2001 Oct 30. J Biol Chem. 2002. PMID: 11687585
-
Bacteriophage T4Dam DNA-(adenine-N(6))-methyltransferase. Comparison of pre-steady state and single turnover methylation of 40-mer duplexes containing two (un)modified target sites.J Biol Chem. 2004 Nov 26;279(48):50012-8. doi: 10.1074/jbc.M409786200. Epub 2004 Sep 16. J Biol Chem. 2004. PMID: 15375160
-
Pre-steady state kinetics of bacteriophage T4 dam DNA-[N(6)-adenine] methyltransferase: interaction with native (GATC) or modified sites.Nucleic Acids Res. 2000 Nov 1;28(21):4207-11. doi: 10.1093/nar/28.21.4207. Nucleic Acids Res. 2000. PMID: 11058118 Free PMC article.
-
[Molecular enzymology of phage T4 Dam DNA-methyltransferase].Mol Biol (Mosk). 2004 Sep-Oct;38(5):869-85. Mol Biol (Mosk). 2004. PMID: 15554189 Review. Russian.
-
Structure and function of DNA methyltransferases.Annu Rev Biophys Biomol Struct. 1995;24:293-318. doi: 10.1146/annurev.bb.24.060195.001453. Annu Rev Biophys Biomol Struct. 1995. PMID: 7663118 Review.
Cited by
-
Structure, function and mechanism of exocyclic DNA methyltransferases.Biochem J. 2006 Oct 15;399(2):177-90. doi: 10.1042/BJ20060854. Biochem J. 2006. PMID: 16987108 Free PMC article. Review.
-
Organization of the BcgI restriction-modification protein for the transfer of one methyl group to DNA.Nucleic Acids Res. 2013 Jan 7;41(1):405-17. doi: 10.1093/nar/gks1000. Epub 2012 Nov 11. Nucleic Acids Res. 2013. PMID: 23147004 Free PMC article.
-
Nucleotide flipping by restriction enzymes analyzed by 2-aminopurine steady-state fluorescence.Nucleic Acids Res. 2007;35(14):4792-9. doi: 10.1093/nar/gkm513. Epub 2007 Jul 7. Nucleic Acids Res. 2007. PMID: 17617640 Free PMC article.
-
Time-resolved fluorescence studies of nucleotide flipping by restriction enzymes.Nucleic Acids Res. 2009 Nov;37(20):6859-70. doi: 10.1093/nar/gkp688. Epub 2009 Sep 8. Nucleic Acids Res. 2009. PMID: 19740769 Free PMC article.
-
Dimerization of the bacterial RsrI N6-adenine DNA methyltransferase.Nucleic Acids Res. 2006 Feb 6;34(3):806-15. doi: 10.1093/nar/gkj486. Print 2006. Nucleic Acids Res. 2006. PMID: 16464821 Free PMC article.
References
-
- Cheng X. (1995) Structure and function of DNA methyltransferases. Annu. Rev. Biophys. Biomol. Struct., 24, 293–318. - PubMed
-
- Wu J.C. and Santi,D.V. (1987) Kinetic and catalytic mechanism of HhaI methyltransferase. J. Biol. Chem., 262, 4778–4786. - PubMed
-
- Cheng X., Kumar,S., Posfai,J., Pflugrath,J.W. and Roberts,R.J. (1993) Crystal structure of the HhaI DNA methyltransferase complexed with S-adenosyl-l-methionine. Cell, 74, 299–307. - PubMed
-
- Cheng X., Kumar,S., Klimasauskas,S. and Roberts,R.J. (1993) Crystal structure of the HhaI DNA methyltransferase. Cold Spring Harbor Symp. Quant. Biol., 58, 331–338. - PubMed
-
- Klimasauskas S., Kumar,S., Roberts,R.J. and Cheng,X. (1994) HhaI methyltransferase flips its target base out of the DNA helix. Cell, 76, 357–369. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

