Hormone-induced nucleosome positioning in the MMTV promoter is reversible
- PMID: 11387213
- PMCID: PMC125491
- DOI: 10.1093/emboj/20.11.2802
Hormone-induced nucleosome positioning in the MMTV promoter is reversible
Abstract
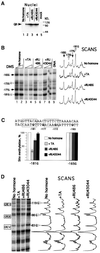
The mouse mammary tumor virus (MMTV) promoter is induced by glucocorticoid hormone via the glucocorticoid receptor (GR). The hormone-triggered effects on MMTV transcription and chromatin structure were studied in Xenopus oocytes. We previously showed that the nucleosomes organizing the MMTV promoter became translationally positioned upon hormone induction. A single GR-binding site was necessary and sufficient for the chromatin events to occur, while transcription and basal promoter elements were dispensable. Here we show that addition of the hormone antagonists RU486 or RU43044 to the previously hormone-induced MMTV promoter results in cessation of transcription and loss of chromatin remodeling and nucleosome positioning. In vivo footprinting demonstrated agonist- and RU486-induced GR binding to its DNA response element (GRE), while the other antagonist, RU43044, did not promote GR-GRE interaction. These results demonstrate that induction and maintenance of nucleosome positioning is an active process that requires constant 'pressure' of agonist-GR-recruited chromatin-modifying factor(s) rather than GR-DNA binding itself.
Figures



References
-
- Agalioti T., Lomvardas,S., Parekh,B., Yie,J., Maniatis,T., Thanos,D., El-Osta,A. and Wolffe,A.P. (2000) Ordered recruitment of chromatin modifying and general transcription factors to the IFN-β promoter. Cell, 103, 667–678. - PubMed
-
- Almouzni G. and Wolffe,A.P. (1993) Replication-coupled chromatin assembly is required for repression of basal transcription in vivo. Genes Dev., 7, 2033–2047. - PubMed
-
- Becker P.B., Gloss,B., Schmid,W., Strahle,U. and Schutz,G. (1986) In vivo protein–DNA interactions in a glucocorticoid response element require the presence of the hormone. Nature, 324, 686–688. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

