Inability to enter S phase and defective RNA polymerase II CTD phosphorylation in mice lacking Mat1
- PMID: 11387217
- PMCID: PMC125252
- DOI: 10.1093/emboj/20.11.2844
Inability to enter S phase and defective RNA polymerase II CTD phosphorylation in mice lacking Mat1
Abstract
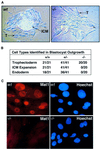
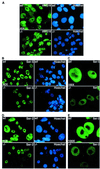
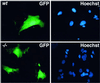
The trimeric Cdk7-cyclin H-Mat1 complex comprises the kinase subunit of basal transcription factor TFIIH and has been shown to function as a cyclin-dependent kinase (Cdk)-activating kinase. Herein we report that disruption of the murine Mat1 gene leads to peri-implantation lethality coincident with depletion of maternal Mat1 protein. In culture, Mat1(-/-) blastocysts gave rise to viable post-mitotic trophoblast giant cells while mitotic lineages failed to proliferate and survive. In contrast to wild-type trophoblast giant cells, Mat1(-/-) cells exhibited a rapid arrest in endoreduplication, which was characterized by an inability to enter S phase. Additionally, Mat1(-/-) cells exhibited defects in phosphorylation of the C-terminal domain (CTD) of RNA polymerase II on both Ser5 and Ser2 of the heptapeptide repeat. Despite this, Mat1(-/-) cells demonstrated apparent transcriptional and translational integrity. These data indicate an essential role for Mat1 in progression through the endocycle and suggest that while Mat1 modulates CTD phosphorylation, it does not appear to be essential for RNA polymerase II-mediated transcription.
Figures





References
-
- Barlow P., Owen,D.A. and Graham,C. (1972) DNA synthesis in the preimplantation mouse embryo. J. Embryol. Exp. Morphol., 27, 431–445. - PubMed
-
- Busso D., Keriel,A., Sandrock,B., Poterszman,A., Gileadi,O. and Egly,J.M. (2000) Distinct regions of MAT1 regulate cdk7 kinase and TFIIH transcription activities. J. Biol. Chem., 275, 22815–22823. - PubMed
-
- Chen D., Riedl,T., Washbrook,E., Pace,P.E., Coombes,R.C., Egly,J.M. and Ali,S. (2000) Activation of estrogen receptor α by S118 phosphorylation involves a ligand-dependent interaction with TFIIH and participation of CDK7. Mol. Cell, 6, 127–137. - PubMed
-
- Dahmus M.E. (1996) Reversible phosphorylation of the C-terminal domain of RNA polymerase II. J. Biol. Chem., 271, 19009–19012. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

