In vitro processing of the 16S rRNA of the thermophilic archaeon Sulfolobus solfataricus
- PMID: 11395449
- PMCID: PMC95268
- DOI: 10.1128/JB.183.13.3866-3874.2001
In vitro processing of the 16S rRNA of the thermophilic archaeon Sulfolobus solfataricus
Abstract
In this paper we have analyzed the processing in vitro of the 16S rRNA of the thermophilic archaeon Sulfolobus solfataricus, using pre-rRNA substrates transcribed in vitro and different protein preparations as the source of processing enzymes. We show that the 5' external transcribed spacer of the S. solfataricus pre-rRNA transcript contains a target site for a specific endonuclease, which recognizes a conserved sequence also existing in the early A0 and 0 processing sites of Saccharomyces cerevisiae and vertebrates. This site is present in other members of the kingdom Crenarchaeota but apparently not in the Euryarchaeota. Furthermore, S. solfataricus pre-16S RNA is processed within the double-helical stem formed by the inverted repeats flanking the 16S RNA sequence, in correspondence with a bulge-helix-bulge motif. The endonuclease responsible for this cleavage is present in both the Crenarchaeota and the Euryarchaeota. The processing pattern remained the same when the substrate was a 30S ribonucleoprotein particle instead of the naked RNA. Maturation of either the 5' or the 3' end of the 16S RNA molecule was not observed, suggesting either that maturation requires conditions not easily reproducible in vitro or that the responsible endonucleases are scarcely represented in cell extracts.
Figures




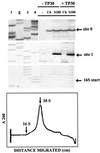

References
-
- Abou Elela S, Igel H, Ares M., Jr RNase III cleaves eukaryotic pre-ribosomal RNA at a U3 snoRNP-dependent site. Cell. 1996;85:115–124. - PubMed
-
- Deng W P, Nickoloff J A. Site-directed mutagenesis of virtually any plasmid by eliminating a unique site. Anal Biochem. 1992;200:81–88. - PubMed
-
- Dennis P P. Multiple promoters for the transcription of the ribosomal RNA gene cluster in Halobacterium cutirubrum. J Mol Biol. 1985;186:457–461. - PubMed
-
- De Rosa M, Gambacorta A, Bu'lock J D. Extremely thermoacidophilic bacteria convergent with Sulfolobus acidocaldarius. J Gen Microbiol. 1975;86:156–164. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

