Localization of a germinant receptor protein (GerBA) to the inner membrane of Bacillus subtilis spores
- PMID: 11395462
- PMCID: PMC95281
- DOI: 10.1128/JB.183.13.3982-3990.2001
Localization of a germinant receptor protein (GerBA) to the inner membrane of Bacillus subtilis spores
Abstract
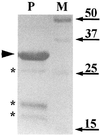
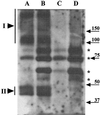
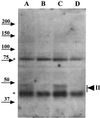
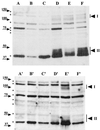
Dormant Bacillus subtilis spores germinate in response to specific nutrients called germinants, which are recognized by multisubunit receptor complexes encoded by members of the gerA family of operons, of which the gerB operon is a member. The germinant receptors are expected to be membrane associated, but there is some debate about whether they are located in the inner or outer spore membrane. In this study we have used Western blot analysis to determine the precise location of GerBA, a gerB-encoded receptor protein, in various spore fractions. GerBA was not extracted from spores by a decoating treatment that removes the coat and outer membrane but was present in lysates from decoated spores and in the insoluble fraction (termed P100) from such lysates that contained inner-membrane vesicles. GerBA was also solubilized from the P100 fraction with detergent but not with high salt. These findings suggest that GerBA is an integral membrane protein located in the spore's inner membrane. Consistent with this idea, GerBA was present in the cell membrane of the outgrowing spore, a membrane that is derived from the dormant spore's inner membrane. Based on these observations we propose that GerBA and probably the entire GerB germinant receptor are located in the inner membrane of the dormant spore. We also estimated that there are only 24 to 40 molecules of GerBA per spore, a number that is consistent with the previously reported low level of gerB operon expression and with the putative receptor function of the proteins encoded by the gerB operon.
Figures

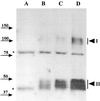
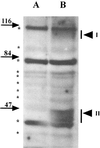
Similar articles
-
Numbers of individual nutrient germinant receptors and other germination proteins in spores of Bacillus subtilis.J Bacteriol. 2013 Aug;195(16):3575-82. doi: 10.1128/JB.00377-13. Epub 2013 Jun 7. J Bacteriol. 2013. PMID: 23749970 Free PMC article.
-
Effects of overexpression of nutrient receptors on germination of spores of Bacillus subtilis.J Bacteriol. 2003 Apr;185(8):2457-64. doi: 10.1128/JB.185.8.2457-2464.2003. J Bacteriol. 2003. PMID: 12670969 Free PMC article.
-
Isolation and characterization of mutations in Bacillus subtilis that allow spore germination in the novel germinant D-alanine.J Bacteriol. 1999 Jun;181(11):3341-50. doi: 10.1128/JB.181.11.3341-3350.1999. J Bacteriol. 1999. PMID: 10348844 Free PMC article.
-
[Overview of study on Bacillus subtilis spores].Yakugaku Zasshi. 2013;133(7):783-97. doi: 10.1248/yakushi.13-00143. Yakugaku Zasshi. 2013. PMID: 23811766 Review. Japanese.
-
Spores of Bacillus subtilis: their resistance to and killing by radiation, heat and chemicals.J Appl Microbiol. 2006 Sep;101(3):514-25. doi: 10.1111/j.1365-2672.2005.02736.x. J Appl Microbiol. 2006. PMID: 16907802 Review.
Cited by
-
Numbers of individual nutrient germinant receptors and other germination proteins in spores of Bacillus subtilis.J Bacteriol. 2013 Aug;195(16):3575-82. doi: 10.1128/JB.00377-13. Epub 2013 Jun 7. J Bacteriol. 2013. PMID: 23749970 Free PMC article.
-
Effects of overexpression of nutrient receptors on germination of spores of Bacillus subtilis.J Bacteriol. 2003 Apr;185(8):2457-64. doi: 10.1128/JB.185.8.2457-2464.2003. J Bacteriol. 2003. PMID: 12670969 Free PMC article.
-
Membrane Proteomes and Ion Transporters in Bacillus anthracis and Bacillus subtilis Dormant and Germinating Spores.J Bacteriol. 2019 Feb 25;201(6):e00662-18. doi: 10.1128/JB.00662-18. Print 2019 Mar 15. J Bacteriol. 2019. PMID: 30602489 Free PMC article.
-
The products of the spoVA operon are involved in dipicolinic acid uptake into developing spores of Bacillus subtilis.J Bacteriol. 2002 Jan;184(2):584-7. doi: 10.1128/JB.184.2.584-587.2002. J Bacteriol. 2002. PMID: 11751839 Free PMC article.
-
A novel lipolytic enzyme, YcsK (LipC), located in the spore coat of Bacillus subtilis, is involved in spore germination.J Bacteriol. 2007 Mar;189(6):2369-75. doi: 10.1128/JB.01527-06. Epub 2007 Jan 12. J Bacteriol. 2007. PMID: 17220230 Free PMC article.
References
-
- Bagyan I, Noback M, Bron S, Paidhungat M, Setlow P. Characterization of yhcN, a new forespore-specific gene of Bacillus subtilis. Gene. 1998;212:179–188. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

