CREB is one component of the binding complex of the Ces-2/E2A-HLF binding element and is an integral part of the interleukin-3 survival signal
- PMID: 11416141
- PMCID: PMC87131
- DOI: 10.1128/MCB.21.14.4636-4646.2001
CREB is one component of the binding complex of the Ces-2/E2A-HLF binding element and is an integral part of the interleukin-3 survival signal
Abstract
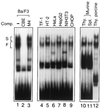
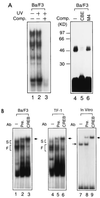
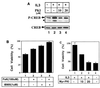
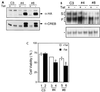
The Ces-2/E2A-HLF binding element (CBE) is recognized by Caenorhabditis elegans death specification gene product Ces-2 and human acute lymphocytic leukemia oncoprotein E2A-HLF. In an attempt to identify a cellular CBE-binding protein(s) that may be involved in apoptosis regulation in mammals, multiple nuclear binding complexes of CBE were identified in various mammalian cell lines and tissues by electrophoretic mobility shift assay. Cyclic AMP (cAMP)-responsive element (CRE)-binding protein (CREB) was present in one major CBE complex of Ba/F3 and TF-1 cells, and both in vitro-translated and Escherichia coli-synthesized CREB bound to CBE. Activation of CREB by cAMP-elevating chemicals or the catalytic subunit of protein kinase A (PKAc) resulted in induction of the CBE-driven reporter gene. Stimulation of Ba/F3 cells with interleukin-3 (IL-3) promptly induced phosphorylation of CREB at serine(133) partially via a PKA-dependent pathway. Consistently, Ba/F3 cell survival in the absence of IL-3 was prolonged by activation of PKA. Conversely, treatment of cells with a PKA inhibitor or expression of the dominant negative forms of the regulatory subunit type I of PKA and CREB overrode the survival activity of IL-3. Last, the bcl-2 gene was demonstrated to be one candidate cellular target of the CREB-containing CBE complex, as mutations in the CRE and CBE sites significantly reduced the IL-3 inducibility of the bcl-2 promoter. Together, our results suggest that CREB is one cellular counterpart of Ces-2/E2A-HLF and is part of IL-3 dependent apoptosis regulation in hematopoietic cells.
Figures


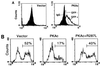

References
-
- Barton K, Muthusamy N, Chanyangam M, Fischer C, Clendenin C, Leiden J M. Defective thymocyte proliferation and IL-2 production in transgenic mice expressing a dominant-negative form of CREB. Nature. 1996;379:81–85. - PubMed
-
- Blendy J A, Kaestner K H, Weinbauer G F, Nieschlag E, Schutz G. Severe impairment of spermatogenesis in mice lacking the CREM gene. Nature. 1996;380:162–165. - PubMed
-
- Chadosh L A. Mobility shift DNA-binding assay using gel electrophoresis. In: Ausubel F M, Brent R, Kingston R E, Moore D D, Seidman J G, Smith J A, Struhl K, editors. Current protocols in molecular biology. New York, N.Y: Wiley Publishers; 1988. p. 12.2.
-
- Chao J R, Wang J M, Lee S F, Peng H W, Lin Y H, Chou C H, Li J C, Huang H M, Chou C K, Kuo M L, Yen J J, Yang-Yen H F. mcl-1 is an immediate-early gene activated by the granulocyte-macrophage colony-stimulating factor (GM-CSF) signaling pathway and is one component of the GM-CSF viability response. Mol Cell Biol. 1998;18:4883–4898. - PMC - PubMed
-
- Chrivia J C, Kwok R P, Lamb N, Hagiwara M, Montminy M R, Goodman R H. Phosphorylated CREB binds specifically to the nuclear protein CBP. Nature. 1993;365:855–859. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous
