The ribonucleotide reductase inhibitor Sml1 is a new target of the Mec1/Rad53 kinase cascade during growth and in response to DNA damage
- PMID: 11432841
- PMCID: PMC125510
- DOI: 10.1093/emboj/20.13.3544
The ribonucleotide reductase inhibitor Sml1 is a new target of the Mec1/Rad53 kinase cascade during growth and in response to DNA damage
Abstract
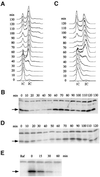
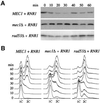
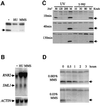
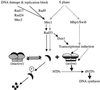
The evolutionarily conserved protein kinases Mec1 and Rad53 are required for checkpoint response and growth. Here we show that their role in growth is to remove the ribonucleotide reductase inhibitor Sml1 to ensure DNA replication. Sml1 protein levels fluctuate during the cell cycle, being lowest during S phase. The disappearance of Sml1 protein in S phase is due to post-transcriptional regulation and is associated with protein phosphorylation. Both phosphorylation and diminution of Sml1 require MEC1 and RAD53. More over, failure to remove Sml1 in mec1 and rad53 mutants results in incomplete DNA replication, defective mitochondrial DNA propagation, decreased dNTP levels and cell death. Interestingly, similar regulation of Sml1 also occurs after DNA damage. In this case, the regulation requires MEC1 and RAD53, as well as other checkpoint genes. Therefore, Sml1 is a new target of the DNA damage checkpoint and its removal is a conserved function of Mec1 and Rad53 during growth and after damage.
Figures




Similar articles
-
The Dun1 checkpoint kinase phosphorylates and regulates the ribonucleotide reductase inhibitor Sml1.Proc Natl Acad Sci U S A. 2002 Mar 19;99(6):3746-51. doi: 10.1073/pnas.062502299. Proc Natl Acad Sci U S A. 2002. PMID: 11904430 Free PMC article.
-
Mutational and structural analyses of the ribonucleotide reductase inhibitor Sml1 define its Rnr1 interaction domain whose inactivation allows suppression of mec1 and rad53 lethality.Mol Cell Biol. 2000 Dec;20(23):9076-83. doi: 10.1128/MCB.20.23.9076-9083.2000. Mol Cell Biol. 2000. PMID: 11074005 Free PMC article.
-
A suppressor of two essential checkpoint genes identifies a novel protein that negatively affects dNTP pools.Mol Cell. 1998 Sep;2(3):329-40. doi: 10.1016/s1097-2765(00)80277-4. Mol Cell. 1998. PMID: 9774971
-
Yeast Dun1 kinase regulates ribonucleotide reductase inhibitor Sml1 in response to iron deficiency.Mol Cell Biol. 2014 Sep;34(17):3259-71. doi: 10.1128/MCB.00472-14. Epub 2014 Jun 23. Mol Cell Biol. 2014. PMID: 24958100 Free PMC article.
-
Inactivation of Ku-mediated end joining suppresses mec1Delta lethality by depleting the ribonucleotide reductase inhibitor Sml1 through a pathway controlled by Tel1 kinase and the Mre11 complex.Mol Cell Biol. 2005 Dec;25(23):10652-64. doi: 10.1128/MCB.25.23.10652-10664.2005. Mol Cell Biol. 2005. PMID: 16287875 Free PMC article.
Cited by
-
Proteome analysis of Saccharomyces cerevisiae after methyl methane sulfonate (MMS) treatment.Biochem Biophys Rep. 2020 Oct 3;24:100820. doi: 10.1016/j.bbrep.2020.100820. eCollection 2020 Dec. Biochem Biophys Rep. 2020. PMID: 33072891 Free PMC article.
-
The checkpoint transcriptional response: make sure to turn it off once you are satisfied.Cell Cycle. 2012 Sep 1;11(17):3166-74. doi: 10.4161/cc.21197. Epub 2012 Aug 16. Cell Cycle. 2012. PMID: 22895177 Free PMC article.
-
Nuclear localization of the Saccharomyces cerevisiae ribonucleotide reductase small subunit requires a karyopherin and a WD40 repeat protein.Proc Natl Acad Sci U S A. 2006 Jan 31;103(5):1422-7. doi: 10.1073/pnas.0510516103. Epub 2006 Jan 23. Proc Natl Acad Sci U S A. 2006. PMID: 16432237 Free PMC article.
-
Tel1 kinase and subtelomere-bound Tbf1 mediate preferential elongation of short telomeres by telomerase in yeast.EMBO Rep. 2007 Nov;8(11):1080-5. doi: 10.1038/sj.embor.7401082. Epub 2007 Oct 5. EMBO Rep. 2007. PMID: 17917674 Free PMC article.
-
The protein kinase Snf1 is required for tolerance to the ribonucleotide reductase inhibitor hydroxyurea.Mol Cell Biol. 2004 Mar;24(6):2560-72. doi: 10.1128/MCB.24.6.2560-2572.2004. Mol Cell Biol. 2004. PMID: 14993292 Free PMC article.
References
-
- Adams A., Gottschling,D.E., Kaiser,C.A. and Stearns,T. (1997) Methods in Yeast Genetics. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY.
-
- Caspari T. (2000) How to activate p53. Curr. Biol., 10, R315–R317. - PubMed
-
- Chabes A., Domkin,V. and Thelander,L. (1999) Yeast Sml1, a protein inhibitor of ribonucleotide reductase. J. Biol. Chem., 274, 36679–36683. - PubMed
-
- Christianson T.W., Sikorski,R.S., Dante,M., Shero,J.H. and Hieter,P. (1992) Multifunctional yeast high-copy-number shuttle vectors. Gene, 110, 119–122. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

