Human immunodeficiency virus type 1 integrase: arrangement of protein domains in active cDNA complexes
- PMID: 11432843
- PMCID: PMC125503
- DOI: 10.1093/emboj/20.13.3565
Human immunodeficiency virus type 1 integrase: arrangement of protein domains in active cDNA complexes
Abstract
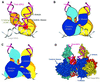
Early steps of retroviral replication involve reverse transcription of the viral RNA genome and integration of the resulting cDNA copy into a chromosome of the host cell. The viral-encoded integrase protein carries out the initial DNA breaking and joining reactions that mediate integration. The organization of the active integrase-DNA complex is unknown, though integrase is known to act as a multimer, and high resolution structures of the isolated integrase domains have been determined. Here we use site-specific cross-linking based on disulfide bond formation to map integrase-DNA contacts in active complexes. We establish that the DNA-binding C-terminal domain of one integrase monomer acts with the central catalytic domain from another monomer at each viral cDNA end. These data allow detailed modeling of an integrase tetramer in which pairs of trans interactions link integrase dimers bound to substrate DNA. We also detected a conformational change in integrase- DNA complexes accompanying cleavage of the viral cDNA terminus.
Figures


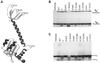

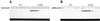

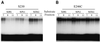
Similar articles
-
Division of labor within human immunodeficiency virus integrase complexes: determinants of catalysis and target DNA capture.J Virol. 2005 Dec;79(24):15376-87. doi: 10.1128/JVI.79.24.15376-15387.2005. J Virol. 2005. PMID: 16306609 Free PMC article.
-
Paired DNA three-way junctions as scaffolds for assembling integrase complexes.Virology. 2001 Aug 1;286(2):304-16. doi: 10.1006/viro.2001.0978. Virology. 2001. PMID: 11485398
-
Crystal structure of an active two-domain derivative of Rous sarcoma virus integrase.J Mol Biol. 2000 Feb 18;296(2):535-48. doi: 10.1006/jmbi.1999.3463. J Mol Biol. 2000. PMID: 10669607
-
Structure and function of HIV-1 integrase.Curr Top Med Chem. 2004;4(9):965-77. doi: 10.2174/1568026043388547. Curr Top Med Chem. 2004. PMID: 15134551 Review.
-
HIV DNA integration.Cold Spring Harb Perspect Med. 2012 Jul;2(7):a006890. doi: 10.1101/cshperspect.a006890. Cold Spring Harb Perspect Med. 2012. PMID: 22762018 Free PMC article. Review.
Cited by
-
A possible role for the asymmetric C-terminal domain dimer of Rous sarcoma virus integrase in viral DNA binding.PLoS One. 2013;8(2):e56892. doi: 10.1371/journal.pone.0056892. Epub 2013 Feb 22. PLoS One. 2013. PMID: 23451105 Free PMC article.
-
Single-particle image reconstruction of a tetramer of HIV integrase bound to DNA.J Mol Biol. 2007 Feb 9;366(1):286-94. doi: 10.1016/j.jmb.2006.11.029. Epub 2006 Nov 11. J Mol Biol. 2007. PMID: 17157316 Free PMC article.
-
Structure-based modeling of the functional HIV-1 intasome and its inhibition.Proc Natl Acad Sci U S A. 2010 Sep 7;107(36):15910-5. doi: 10.1073/pnas.1002346107. Epub 2010 Aug 23. Proc Natl Acad Sci U S A. 2010. PMID: 20733078 Free PMC article.
-
Genetic analyses of DNA-binding mutants in the catalytic core domain of human immunodeficiency virus type 1 integrase.J Virol. 2005 Feb;79(4):2493-505. doi: 10.1128/JVI.79.4.2493-2505.2005. J Virol. 2005. PMID: 15681450 Free PMC article.
-
Subunit-specific protein footprinting reveals significant structural rearrangements and a role for N-terminal Lys-14 of HIV-1 Integrase during viral DNA binding.J Biol Chem. 2008 Feb 29;283(9):5632-41. doi: 10.1074/jbc.M705241200. Epub 2007 Dec 19. J Biol Chem. 2008. PMID: 18093980 Free PMC article.
References
-
- Aldaz H., Schuster,E. and Baker,T.A. (1996) The interwoven architecture of the Mu transposase couples DNA synapsis to catalysis. Cell, 85, 257–269. - PubMed
-
- Asante-Appiah E., Seeholzer,S.H. and Skalka,A.M. (1998) Structural determinants of metal-induced conformational changes in HIV-1 integrase. J. Biol. Chem., 273, 35078–35087. - PubMed
-
- Bujacz G., Jaskolski,M., Alexandratos,J., Wlodawer,A., Merkel,G., Katz,R.A. and Skalka,A.M. (1995) High-resolution structure of the catalytic domain of avian sarcoma virus integrase. J. Mol. Biol., 253, 333–346. - PubMed
-
- Bushman F.D., Fujiwara,T. and Craigie,R. (1990) Retroviral DNA integration directed by HIV integration protein in vitro. Science, 249, 1555–1558. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

