Quantification of splice variants using real-time PCR
- PMID: 11433044
- PMCID: PMC55792
- DOI: 10.1093/nar/29.13.e68
Quantification of splice variants using real-time PCR
Abstract
A reliable and robust method for measuring the expression of alternatively spliced transcripts is an important step in investigating the significance of each variant. So far, accurate quantification of splice variants has been laborious and difficult due to the intrinsic limitations of conventional methods. The many advantages of real-time PCR have made this technique attractive to study its application in quantification of splice isoforms. We use skipping of exon 37 in the NF1 gene as a model to compare and evaluate the different strategies for quantitating splice variants using real-time PCR. An overview of three different possibilities for detecting alternative transcripts is given. We propose the use of a boundary-spanning primer to quantify isoforms that differ greatly in abundance. We describe here a novel method for creating a reliable standard curve using one plasmid containing both alternative transcripts. In addition, we validate the use of an absolute standard curve based on a dilution series of fluorometrically quantified PCR products.
Figures

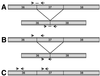



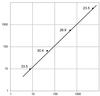
References
-
- Sambrook J., Fritsch,E.F. and Maniatis,T. (1989) Molecular Cloning: A laboratory Manual, 2nd Edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, NY.
-
- Hod Y. (1992) A simplified ribonuclease protection assay. Biotechniques, 13, 852–854. - PubMed
-
- Saccomanno C.F., Bordonaro,M., Chen,J.S. and Nordstrom,J.L. (1992) A faster ribonuclease protection assay. Biotechniques, 13, 846–850. - PubMed
-
- Ferré F. Quantitative or semi-quantitative PCR: reality versus myth. (1992) PCR Methods Appl., 2, 1–9. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

