DNA binding by Kaposi's sarcoma-associated herpesvirus lytic switch protein is necessary for transcriptional activation of two viral delayed early promoters
- PMID: 11435557
- PMCID: PMC114405
- DOI: 10.1128/JVI.75.15.6786-6799.2001
DNA binding by Kaposi's sarcoma-associated herpesvirus lytic switch protein is necessary for transcriptional activation of two viral delayed early promoters
Abstract
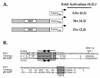
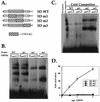
Kaposi's sarcoma-associated herpesvirus (KSHV; also known as human herpesvirus-8) establishes latent and lytic infections in both lymphoid and endothelial cells and has been associated with diseases of both cell types. The KSHV open reading frame 50 (ORF50) protein is a transcriptional activator that plays a central role in the reactivation of lytic viral replication from latency. Here we identify and characterize a DNA binding site for the ORF50 protein that is shared by the promoters of two delayed early genes (ORF57 and K-bZIP). Transfer of this element to heterologous promoters confers on them high-level responsiveness to ORF50, indicating that the element is both necessary and sufficient for activation. The element consists of a conserved 12-bp palindromic sequence and less conserved sequences immediately 3' to it. Mutational analysis reveals that sequences within the palindrome are critical for binding and activation by ORF50, but the presence of a palindrome itself is not absolutely required. The 3' flanking sequences also play a critical role in DNA binding and transactivation. The strong concordance of DNA binding in vitro with transcriptional activation in vivo strongly implies that sequence-specific DNA binding is necessary for ORF50-mediated activation through this element. Expression of truncated versions of the ORF50 protein reveals that DNA binding is mediated by the amino-terminal 272 amino acids of the polypeptide.
Figures



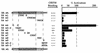



Similar articles
-
Direct interactions of Kaposi's sarcoma-associated herpesvirus/human herpesvirus 8 ORF50/Rta protein with the cellular protein octamer-1 and DNA are critical for specifying transactivation of a delayed-early promoter and stimulating viral reactivation.J Virol. 2007 Aug;81(16):8451-67. doi: 10.1128/JVI.00265-07. Epub 2007 May 30. J Virol. 2007. PMID: 17537858 Free PMC article.
-
Characterization of Kaposi's sarcoma-associated herpesvirus (KSHV) K1 promoter activation by Rta.Virology. 2006 May 10;348(2):309-27. doi: 10.1016/j.virol.2006.02.007. Epub 2006 Mar 20. Virology. 2006. PMID: 16546233
-
Amplification of the Kaposi's sarcoma-associated herpesvirus/human herpesvirus 8 lytic origin of DNA replication is dependent upon a cis-acting AT-rich region and an ORF50 response element and the trans-acting factors ORF50 (K-Rta) and K8 (K-bZIP).Virology. 2004 Jan 20;318(2):542-55. doi: 10.1016/j.virol.2003.10.016. Virology. 2004. PMID: 14972523
-
Lytic cycle switches of oncogenic human gammaherpesviruses.Adv Cancer Res. 2007;97:81-109. doi: 10.1016/S0065-230X(06)97004-3. Adv Cancer Res. 2007. PMID: 17419942 Review.
-
Kaposi's sarcoma-associated herpesvirus immediate early gene activity.Front Biosci. 2004 Sep 1;9:2245-72. doi: 10.2741/1394. Front Biosci. 2004. PMID: 15353285 Review.
Cited by
-
Murine Gammaherpesvirus 68 ORF48 Is an RTA-Responsive Gene Product and Functions in both Viral Lytic Replication and Latency during In Vivo Infection.J Virol. 2015 Jun;89(11):5788-800. doi: 10.1128/JVI.00406-15. Epub 2015 Mar 11. J Virol. 2015. PMID: 25762743 Free PMC article.
-
Role of CCAAT/enhancer-binding protein alpha (C/EBPalpha) in activation of the Kaposi's sarcoma-associated herpesvirus (KSHV) lytic-cycle replication-associated protein (RAP) promoter in cooperation with the KSHV replication and transcription activator (RTA) and RAP.J Virol. 2003 Jan;77(1):600-23. doi: 10.1128/jvi.77.1.600-623.2003. J Virol. 2003. PMID: 12477864 Free PMC article.
-
Kaposi's sarcoma-associated herpesvirus/human herpesvirus 8 RTA reactivates murine gammaherpesvirus 68 from latency.J Virol. 2005 Mar;79(5):3217-22. doi: 10.1128/JVI.79.5.3217-3222.2005. J Virol. 2005. PMID: 15709045 Free PMC article.
-
A herpesvirus transactivator and cellular POU proteins extensively regulate DNA binding of the host Notch signaling protein RBP-Jκ to the virus genome.J Biol Chem. 2019 Aug 30;294(35):13073-13092. doi: 10.1074/jbc.RA118.007331. Epub 2019 Jul 15. J Biol Chem. 2019. PMID: 31308175 Free PMC article.
-
The cellular Notch1 protein promotes KSHV reactivation in an Rta-dependent manner.J Virol. 2024 Aug 20;98(8):e0078824. doi: 10.1128/jvi.00788-24. Epub 2024 Jul 8. J Virol. 2024. PMID: 38975769 Free PMC article.
References
-
- Ambroziak J, Blackbourn D, Herndier B, Glogan R, Gullet J, McDonald A, Lennette E, Levy J. Herpesvirus-like sequences in HIV-infected and uninfected Kaposi's sarcoma patients. Science. 1995;268:582–583. - PubMed
-
- Boshoff C, Schulz T F, Kennedy M M, Graham A K, Fisher C, Thomas A, McGee J O, Weiss R A, O'Leary J J. Kaposi's sarcoma-associated herpesvirus infects endothelial and spindle cells. Nat Med. 1995;1:1274–1278. - PubMed
-
- Boyer T G, Berk A J. Functional interaction of adenovirus E1A with holo-TFIID. Genes Dev. 1993;7:1810–1823. - PubMed
-
- Cesarman E, Chang Y, Moore P S, Said J W, Knowles D M. Kaposi's sarcoma-associated herpesvirus-like DNA sequences in AIDS-related body-cavity-based lymphomas. N Engl J Med. 1995;332:1186–1191. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

