Molecular cloning and sequence analysis of the complestatin biosynthetic gene cluster
- PMID: 11447274
- PMCID: PMC37473
- DOI: 10.1073/pnas.151246498
Molecular cloning and sequence analysis of the complestatin biosynthetic gene cluster
Abstract
Streptomyces lavendulae produces complestatin, a cyclic peptide natural product that antagonizes pharmacologically relevant protein-protein interactions including formation of the C4b,2b complex in the complement cascade and gp120-CD4 binding in the HIV life cycle. Complestatin, a member of the vancomycin group of natural products, consists of an alpha-ketoacyl hexapeptide backbone modified by oxidative phenolic couplings and halogenations. The entire complestatin biosynthetic and regulatory gene cluster spanning ca. 50 kb was cloned and sequenced. It consisted of 16 ORFs, encoding proteins homologous to nonribosomal peptide synthetases, cytochrome P450-related oxidases, ferredoxins, nonheme halogenases, four enzymes involved in 4-hydroxyphenylglycine (Hpg) biosynthesis, transcriptional regulators, and ABC transporters. The nonribosomal peptide synthetase consisted of a priming module, six extending modules, and a terminal thioesterase; their arrangement and domain content was entirely consistent with functions required for the biosynthesis of a heptapeptide or alpha-ketoacyl hexapeptide backbone. Two oxidase genes were proposed to be responsible for the construction of the unique aryl-ether-aryl-aryl linkage on the linear heptapeptide intermediate. Hpg, 3,5-dichloro-Hpg, and 3,5-dichloro-hydroxybenzoylformate are unusual building blocks that repesent five of the seven requisite monomers in the complestatin peptide. Heterologous expression and biochemical analysis of 4-hydroxyphenylglycine transaminon confirmed its role as an aminotransferase responsible for formation of all three precursors. The close similarity but functional divergence between complestatin and chloroeremomycin biosynthetic genes also presents a unique opportunity for the construction of hybrid vancomycin-type antibiotics.
Figures
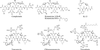



Similar articles
-
Generation of New Complestatin Analogues by Heterologous Expression of the Complestatin Biosynthetic Gene Cluster from Streptomyces chartreusis AN1542.Chembiochem. 2016 Sep 15;17(18):1725-31. doi: 10.1002/cbic.201600241. Epub 2016 Aug 2. Chembiochem. 2016. PMID: 27383040
-
Characterization of biosynthetic gene cluster for the production of virginiamycin M, a streptogramin type A antibiotic, in Streptomyces virginiae.Gene. 2007 May 15;393(1-2):31-42. doi: 10.1016/j.gene.2006.12.035. Epub 2007 Jan 20. Gene. 2007. PMID: 17350183
-
Nonribosomal biosynthesis of vancomycin-type antibiotics: a heptapeptide backbone and eight peptide synthetase modules.Microbiology (Reading). 2002 Apr;148(Pt 4):1105-1118. doi: 10.1099/00221287-148-4-1105. Microbiology (Reading). 2002. PMID: 11932455
-
The enduracidin biosynthetic gene cluster from Streptomyces fungicidicus.Microbiology (Reading). 2006 Oct;152(Pt 10):2969-2983. doi: 10.1099/mic.0.29043-0. Microbiology (Reading). 2006. PMID: 17005978
-
Access to macrocycles with an endo aryl ether and an endo aryl-aryl bond, development and application.Chimia (Aarau). 2011;65(3):168-74. doi: 10.2533/chimia.2011.168. Chimia (Aarau). 2011. PMID: 21528652 Review.
Cited by
-
Altering glycopeptide antibiotic biosynthesis through mutasynthesis allows incorporation of fluorinated phenylglycine residues.RSC Chem Biol. 2024 Aug 12;5(10):1017-34. doi: 10.1039/d4cb00140k. Online ahead of print. RSC Chem Biol. 2024. PMID: 39247680 Free PMC article.
-
Bacterial MbtH-like Proteins Stimulate Nonribosomal Peptide Synthetase-Derived Secondary Metabolism in Filamentous Fungi.ACS Synth Biol. 2019 Aug 16;8(8):1776-1787. doi: 10.1021/acssynbio.9b00106. Epub 2019 Jul 16. ACS Synth Biol. 2019. PMID: 31284717 Free PMC article.
-
Functional Cross-Talk of MbtH-Like Proteins During Thaxtomin Biosynthesis in the Potato Common Scab Pathogen Streptomyces scabiei.Front Microbiol. 2020 Oct 15;11:585456. doi: 10.3389/fmicb.2020.585456. eCollection 2020. Front Microbiol. 2020. PMID: 33178168 Free PMC article.
-
Glycopeptide Antibiotic Resistance Genes: Distribution and Function in the Producer Actinomycetes.Front Microbiol. 2020 Jun 17;11:1173. doi: 10.3389/fmicb.2020.01173. eCollection 2020. Front Microbiol. 2020. PMID: 32655512 Free PMC article. Review.
-
Comparative analysis and insights into the evolution of gene clusters for glycopeptide antibiotic biosynthesis.Mol Genet Genomics. 2005 Aug;274(1):40-50. doi: 10.1007/s00438-005-1156-3. Epub 2005 Jul 9. Mol Genet Genomics. 2005. PMID: 16007453
References
-
- Marahiel M A, Stachelhaus T, Mootz H D. Chem Rev. 1997;97:2651–2673. - PubMed
-
- von Döhren H, Keller U, Vater J, Zocher R. Chem Rev. 1997;97:2675–2705. - PubMed
-
- Weber G, Schörgendorfer K, Schneider-Scherzer E, Leitner E. Curr Genet. 1994;26:120–125. - PubMed
-
- Du L, Chen M, Sanchez C, Shen B. FEMS Microbiol Lett. 2000;189:171–175. - PubMed
-
- Wageningen A M A, Kirkpatrick P N, Williams D H, Harris B R, Kershaw J K, Lennard N J, Jones M, Jones S J M, Solenberg P J. Chem Biol. 1998;5:155–162. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

