Nucleolar protein Nop12p participates in synthesis of 25S rRNA in Saccharomyces cerevisiae
- PMID: 11452019
- PMCID: PMC55798
- DOI: 10.1093/nar/29.14.2938
Nucleolar protein Nop12p participates in synthesis of 25S rRNA in Saccharomyces cerevisiae
Abstract
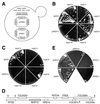
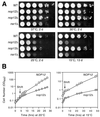
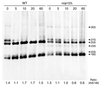
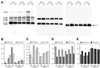
A genetic screen for mutations synthetically lethal with temperature sensitive alleles of nop2 led to the identification of the nucleolar proteins Nop12p and Nop13p in Saccharomyces cerevisiae. NOP12 was identified by complementation of a synthetic lethal growth phenotype in strain YKW35, which contains a single nonsense mutation at codon 359 in an allele termed nop12-1. Database mining revealed that Nop12p was similar to a related protein, Nop13p. Nop12p and Nop13p are not essential for growth and each contains a single canonical RNA recognition motif (RRM). Both share sequence similarity with Nsr1p, a previously identified, non-essential, RRM-containing nucleolar protein. Likely orthologs of Nop12p were identified in Drosophila and Schizosaccharomyces pombe. Deletion of NOP12 resulted in a cold sensitive (cs) growth phenotype at 15 degrees C and slow growth at 20 and 25 degrees C. Growth of a nop12Delta strain at 15 and 20 degrees C resulted in impaired synthesis of 25S rRNA, but not 18S rRNA. A nop13 null strain did not produce an observable growth phenotype under the laboratory conditions examined. Epitope-tagged Nop12p, which complements the cs growth phenotype and restores normal 25S rRNA levels, was localized to the nucleolus by immunofluorescence microscopy. Epitope-tagged Nop13p was distributed primarily in the nucleolus, with a lesser portion localizing to the nucleoplasm. Thus, Nop12p is a novel nucleolar protein required for pre-25S rRNA processing and normal rates of cell growth at low temperatures.
Figures




Similar articles
-
Temperature sensitive nop2 alleles defective in synthesis of 25S rRNA and large ribosomal subunits in Saccharomyces cerevisiae.Nucleic Acids Res. 2001 Jul 15;29(14):2927-37. doi: 10.1093/nar/29.14.2927. Nucleic Acids Res. 2001. PMID: 11452018 Free PMC article.
-
Synthetic lethality with fibrillarin identifies NOP77p, a nucleolar protein required for pre-rRNA processing and modification.EMBO J. 1994 Jul 1;13(13):3136-48. doi: 10.1002/j.1460-2075.1994.tb06612.x. EMBO J. 1994. PMID: 8039506 Free PMC article.
-
The yeast NOP4 gene product is an essential nucleolar protein required for pre-rRNA processing and accumulation of 60S ribosomal subunits.EMBO J. 1994 Jul 1;13(13):3127-35. doi: 10.1002/j.1460-2075.1994.tb06611.x. EMBO J. 1994. PMID: 8039505 Free PMC article.
-
Spb1p is a yeast nucleolar protein associated with Nop1p and Nop58p that is able to bind S-adenosyl-L-methionine in vitro.Mol Cell Biol. 2000 Feb;20(4):1370-81. doi: 10.1128/MCB.20.4.1370-1381.2000. Mol Cell Biol. 2000. PMID: 10648622 Free PMC article.
-
A U3 snoRNP protein with homology to splicing factor PRP4 and G beta domains is required for ribosomal RNA processing.EMBO J. 1993 Jun;12(6):2549-58. doi: 10.1002/j.1460-2075.1993.tb05910.x. EMBO J. 1993. PMID: 8508778 Free PMC article.
Cited by
-
Cysteine of sequence motif VI is essential for nucleophilic catalysis by yeast tRNA m5C methyltransferase.RNA. 2007 Jul;13(7):967-73. doi: 10.1261/rna.515707. Epub 2007 May 2. RNA. 2007. PMID: 17475914 Free PMC article.
-
The eukaryote-specific N-terminal extension of ribosomal protein S31 contributes to the assembly and function of 40S ribosomal subunits.Nucleic Acids Res. 2016 Sep 19;44(16):7777-91. doi: 10.1093/nar/gkw641. Epub 2016 Jul 15. Nucleic Acids Res. 2016. PMID: 27422873 Free PMC article.
-
Temperature sensitive nop2 alleles defective in synthesis of 25S rRNA and large ribosomal subunits in Saccharomyces cerevisiae.Nucleic Acids Res. 2001 Jul 15;29(14):2927-37. doi: 10.1093/nar/29.14.2927. Nucleic Acids Res. 2001. PMID: 11452018 Free PMC article.
-
Ribosome biogenesis in the yeast Saccharomyces cerevisiae.Genetics. 2013 Nov;195(3):643-81. doi: 10.1534/genetics.113.153197. Genetics. 2013. PMID: 24190922 Free PMC article. Review.
-
The assembly factor Erb1 functions in multiple remodeling events during 60S ribosomal subunit assembly in S. cerevisiae.Nucleic Acids Res. 2017 May 5;45(8):4853-4865. doi: 10.1093/nar/gkw1361. Nucleic Acids Res. 2017. PMID: 28115637 Free PMC article.
References
-
- Warner J.R. (1999) The economics of ribosome biosynthesis in yeast. Trends Biochem. Sci., 24, 437–440. - PubMed
-
- Cate J.H., Yusupov,M.M., Yusupova,G.Z., Earnest,T.N. and Noller,H.F. (1999) X-ray crystal structures of 70S ribosome functional complexes. Science, 285, 2095–2104. - PubMed
-
- Venema J. and Tollervey,D. (1995) Processing of pre-ribosomal RNA in Saccharomyces cerevisiae. Yeast, 11, 1629–1650. - PubMed
-
- de la Cruz J., Kressler,D. and Linder,P. (1999) Unwinding RNA in Saccharomyces cerevisiae: DEAD-box proteins and related families. Trends Biochem. Sci., 24, 192–198. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

