Two recombination-dependent DNA replication pathways of bacteriophage T4, and their roles in mutagenesis and horizontal gene transfer
- PMID: 11459968
- PMCID: PMC37436
- DOI: 10.1073/pnas.131007398
Two recombination-dependent DNA replication pathways of bacteriophage T4, and their roles in mutagenesis and horizontal gene transfer
Abstract
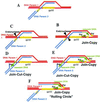
Two major pathways of recombination-dependent DNA replication, "join-copy" and "join-cut-copy," can be distinguished in phage T4: join-copy requires only early and middle genes, but two late proteins, endonuclease VII and terminase, are uniquely important in the join-cut-copy pathway. In wild-type T4, timing of these pathways is integrated with the developmental program and related to transcription and packaging of DNA. In primase mutants, which are defective in origin-dependent lagging-strand DNA synthesis, the late pathway can bypass the lack of primers for lagging-strand DNA synthesis. The exquisitely regulated synthesis of endo VII, and of two proteins from its gene, explains the delay of recombination-dependent DNA replication in primase (as well as topoisomerase) mutants, and the temperature-dependence of the delay. Other proteins (e.g., the single-stranded DNA binding protein and the products of genes 46 and 47) are important in all recombination pathways, but they interact differently with other proteins in different pathways. These homologous recombination pathways contribute to evolution because they facilitate acquisition of any foreign DNA with limited sequence homology during horizontal gene transfer, without requiring transposition or site-specific recombination functions. Partial heteroduplex repair can generate what appears to be multiple mutations from a single recombinational intermediate. The resulting sequence divergence generates barriers to formation of viable recombinants. The multiple sequence changes can also lead to erroneous estimates in phylogenetic analyses.
Figures



Similar articles
-
Recombination and recombination-dependent DNA replication in bacteriophage T4.Annu Rev Genet. 1998;32:379-413. doi: 10.1146/annurev.genet.32.1.379. Annu Rev Genet. 1998. PMID: 9928485 Review.
-
Bypass of a primase requirement for bacteriophage T4 DNA replication in vivo by a recombination enzyme, endonuclease VII.New Biol. 1991 Dec;3(12):1195-205. New Biol. 1991. PMID: 1812964
-
A species barrier between bacteriophages T2 and T4: exclusion, join-copy and join-cut-copy recombination and mutagenesis in the dCTPase genes.Genetics. 1998 Apr;148(4):1461-73. doi: 10.1093/genetics/148.4.1461. Genetics. 1998. PMID: 9560366 Free PMC article.
-
Recombination-dependent DNA replication stimulated by double-strand breaks in bacteriophage T4.J Bacteriol. 1995 Dec;177(23):6844-53. doi: 10.1128/jb.177.23.6844-6853.1995. J Bacteriol. 1995. PMID: 7592477 Free PMC article.
-
Multiple initiation mechanisms adapt phage T4 DNA replication to physiological changes during T4's development.FEMS Microbiol Rev. 1995 Aug;17(1-2):83-98. doi: 10.1111/j.1574-6976.1995.tb00190.x. FEMS Microbiol Rev. 1995. PMID: 7669352 Review.
Cited by
-
Twist and Turn-Topoisomerase Functions in Mitochondrial DNA Maintenance.Int J Mol Sci. 2019 Apr 25;20(8):2041. doi: 10.3390/ijms20082041. Int J Mol Sci. 2019. PMID: 31027213 Free PMC article. Review.
-
Isolation and Characterization of a Novel Phage Collection against Avian-Pathogenic Escherichia coli.Microbiol Spectr. 2023 Jun 15;11(3):e0429622. doi: 10.1128/spectrum.04296-22. Epub 2023 May 4. Microbiol Spectr. 2023. PMID: 37140373 Free PMC article.
-
A short carboxyl-terminal tail is required for single-stranded DNA binding, higher-order structural organization, and stability of the mitochondrial single-stranded annealing protein Mgm101.Mol Biol Cell. 2013 May;24(10):1507-18. doi: 10.1091/mbc.E13-01-0006. Epub 2013 Mar 27. Mol Biol Cell. 2013. PMID: 23536705 Free PMC article.
-
Functional Analysis of the Bacteriophage T4 Rad50 Homolog (gp46) Coiled-coil Domain.J Biol Chem. 2015 Sep 25;290(39):23905-15. doi: 10.1074/jbc.M115.675132. Epub 2015 Aug 4. J Biol Chem. 2015. PMID: 26242734 Free PMC article.
-
Genome annotation and intraviral interactome for the Streptococcus pneumoniae virulent phage Dp-1.J Bacteriol. 2011 Jan;193(2):551-62. doi: 10.1128/JB.01117-10. Epub 2010 Nov 19. J Bacteriol. 2011. PMID: 21097633 Free PMC article.
References
-
- Epstein R H, Bolle A, Steinberg C, Kellenberger E, Boy de la Tour E, Chevalley R, Edgar R, Susman M, Denhardt C, Lielausis I. Cold Spring Harbor Symp Quant Biol. 1964;28:375–392.
-
- Nossal N G. In: Molecular Biology of Bacteriophage T4. Karam J, Drake J W, Kreuzer K N, Mosig G, Hall D H, Eiserling F A, Black L W, Spicer E K, Kutter E, Carlson K, et al., editors. Washington, DC: Am. Soc. Microbiol.; 1994. pp. 43–53.
-
- Alberts B M, Barry J, Bedinger B P, Burke R L, Hibner U, Liu C C, Sheridan R. UCLA Symp Mol Cell Biol. 1980;19:449–473.
-
- Streisinger G. In: Phage and the Origins of Molecular Biology. Cairns J, Stent G S, Watson J D, editors. Plainview, NY: Cold Spring Harbor Lab. Press; 1966. pp. 335–340.
-
- Mosig G. Cold Spring Harbor Symp Quant Biol. 1964;28:35–42.
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

