p63 Gene mutations in eec syndrome, limb-mammary syndrome, and isolated split hand-split foot malformation suggest a genotype-phenotype correlation
- PMID: 11462173
- PMCID: PMC1235479
- DOI: 10.1086/323123
p63 Gene mutations in eec syndrome, limb-mammary syndrome, and isolated split hand-split foot malformation suggest a genotype-phenotype correlation
Abstract
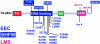
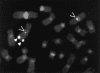
p63 mutations have been associated with EEC syndrome (ectrodactyly, ectodermal dysplasia, and cleft lip/palate), as well as with nonsyndromic split hand-split foot malformation (SHFM). We performed p63 mutation analysis in a sample of 43 individuals and families affected with EEC syndrome, in 35 individuals affected with SHFM, and in three families with the EEC-like condition limb-mammary syndrome (LMS), which is characterized by ectrodactyly, cleft palate, and mammary-gland abnormalities. The results differed for these three conditions. p63 gene mutations were detected in almost all (40/43) individuals affected with EEC syndrome. Apart from a frameshift mutation in exon 13, all other EEC mutations were missense, predominantly involving codons 204, 227, 279, 280, and 304. In contrast, p63 mutations were detected in only a small proportion (4/35) of patients with isolated SHFM. p63 mutations in SHFM included three novel mutations: a missense mutation (K193E), a nonsense mutation (Q634X), and a mutation in the 3' splice site for exon 5. The fourth SHFM mutation (R280H) in this series was also found in a patient with classical EEC syndrome, suggesting partial overlap between the EEC and SHFM mutational spectra. The original family with LMS (van Bokhoven et al. 1999) had no detectable p63 mutation, although it clearly localizes to the p63 locus in 3q27. In two other small kindreds affected with LMS, frameshift mutations were detected in exons 13 and 14, respectively. The combined data show that p63 is the major gene for EEC syndrome, and that it makes a modest contribution to SHFM. There appears to be a genotype-phenotype correlation, in that there is a specific pattern of missense mutations in EEC syndrome that are not generally found in SHFM or LMS.
Figures

References
Electronic-Database Information
-
- GenBank, http://www.ncbi.nlm.nih.gov/Genbank/index.html (for p63 gene sequences [accession numbers AF075430 and AF091627])
-
- IARC p53 Mutation Database, http://www.iarc.fr/p53/index.html (for the distrubution and frequencies of mutations in the p53 gene)
-
- Online Mendelian Inheritance in Man (OMIM), http://www.ncbi.nlm.nih.gov/Omim/ (for SHFM [MIM 183600, 600095, and 313350], and EEC [MIM 129900 and 602077])
References
-
- Brown TC, Jiricny J (1987) A specific mismatch repair event protects mammalian cells from loss of 5-methylcytosine. Cell 50:945–950 - PubMed
-
- Celli J, Duif P, Hamel BCJ, Bamshad M, Kramer B, Smits APT, Newbury-Ecob R, Hennekam RC, Van Buggenhout G, van Haeringen A, Woods CG, van Essen AJ, de Waal R, Vriend G, Haber DA, Yang A, McKeon F, Brunner HG, van Bokhoven H (1999) Heterozygous germline mutations in the p53 homolog p63 are the cause of EEC syndrome. Cell 99:143–153 - PubMed
-
- Chitayat D, Babul R, Silver MM, Jay V, Teshima IE, Babyn P, Becker LE (1996) Terminal deletion of the long arm of chromosome 3 [46,XX,del(3)(q27→qter)]. Am J Med Genet 61:45–48 - PubMed
-
- Church DM, Stotler CJ, Rutter JL, Murrell JR, Trofatter JA, Buckler AJ (1994) Isolation of genes from complex sources of mammalian genomic DNA using exon amplification. Nat Genet 6:98–105 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

