Genetic variation among hospital isolates of methicillin-sensitive Staphylococcus aureus: evidence for horizontal transfer of virulence genes
- PMID: 11473989
- PMCID: PMC88236
- DOI: 10.1128/JCM.39.8.2760-2767.2001
Genetic variation among hospital isolates of methicillin-sensitive Staphylococcus aureus: evidence for horizontal transfer of virulence genes
Abstract
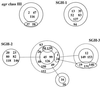
Staphylococcus aureus strains often carry in their genomes virulence genes that are not found in all strains and that may be carried on discrete genetic elements. Strains also differ in that they carry one of four classes of an accessory gene regulator (agr) locus, an operon that regulates virulence factor expression and that has been proposed to be a therapeutic target. To look at their distribution among hospital strains, we investigated 38 methicillin-sensitive S. aureus isolates, classifying the isolates by agr class and screening them for the presence and restriction fragment length polymorphisms (RFLPs) of 12 core and 14 accessory virulence genes. Twenty-three (61%) were agr class I, 10 (26%) were agr class II, and 5 (13%) were agr class III. None were agr class IV. The S. aureus strains had distinguishable RFLP profiles, although clusters of isolates with clearly related core gene profiles were found among our strains, including all five agr class III strains, two sets of six strains within agr class I, and six strains within agr class II. Within these clusters there was evidence of horizontal acquisition and/or loss of multiple accessory virulence genes. Furthermore, two isolates from the same patient were identical except for the presence of the sea gene, indicating that movement of mobile elements may occur in vivo. Several strong correlations with the carriage of virulence genes between strains were seen, including a positive correlation between tst and agr class III and negative correlations between tst and lukE-splB and between lukE-splB and seg-sei. This suggests that the core genome or the presence of accessory genetic elements within a strain may influence acquisition and loss of other elements encoding virulence genes.
Figures

Similar articles
-
Associations between enterotoxin gene cluster types egc1, egc2 and egc3, agr types, enterotoxin and enterotoxin-like gene profiles, and molecular typing characteristics of human nasal carriage and animal isolates of Staphylococcus aureus.J Med Microbiol. 2009 Jan;58(Pt 1):13-25. doi: 10.1099/jmm.0.005215-0. J Med Microbiol. 2009. PMID: 19074649
-
Molecular typing of nasal carriage isolates of Staphylococcus aureus from an Irish university student population based on toxin gene PCR, agr locus types and multiple locus, variable number tandem repeat analysis.J Med Microbiol. 2008 Mar;57(Pt 3):348-358. doi: 10.1099/jmm.0.47734-0. J Med Microbiol. 2008. PMID: 18287299
-
Analysis of the genetic variability of genes encoding the RNA III-activating components Agr and TRAP in a population of Staphylococcus aureus strains isolated from cows with mastitis.J Clin Microbiol. 2002 Nov;40(11):4060-7. doi: 10.1128/JCM.40.11.4060-4067.2002. J Clin Microbiol. 2002. PMID: 12409375 Free PMC article.
-
What role does the quorum-sensing accessory gene regulator system play during Staphylococcus aureus bacteremia?Trends Microbiol. 2014 Dec;22(12):676-85. doi: 10.1016/j.tim.2014.09.002. Epub 2014 Oct 6. Trends Microbiol. 2014. PMID: 25300477 Review.
-
Quorum sensing-mediated regulation of staphylococcal virulence and antibiotic resistance.Future Microbiol. 2014;9(5):669-81. doi: 10.2217/fmb.14.31. Future Microbiol. 2014. PMID: 24957093 Review.
Cited by
-
Virulence characteristic and MLST-agr genetic background of high-level mupirocin-resistant, MRSA isolates from Shanghai and Wenzhou, China.PLoS One. 2012;7(5):e37005. doi: 10.1371/journal.pone.0037005. Epub 2012 May 18. PLoS One. 2012. PMID: 22623969 Free PMC article.
-
Molecular Epidemiology of Staphylococcus aureus Bacteremia: Association of Molecular Factors With the Source of Infection.Front Microbiol. 2018 Sep 25;9:2210. doi: 10.3389/fmicb.2018.02210. eCollection 2018. Front Microbiol. 2018. PMID: 30319561 Free PMC article.
-
Difference in virulence between Staphylococcus aureus isolates causing gangrenous mastitis versus subclinical mastitis in a dairy sheep flock.Vet Res. 2009 Nov-Dec;40(6):56. doi: 10.1051/vetres/2009039. Epub 2009 Jul 7. Vet Res. 2009. PMID: 19576164 Free PMC article.
-
Using Whole Genome Analysis to Examine Recombination across Diverse Sequence Types of Staphylococcus aureus.PLoS One. 2015 Jul 10;10(7):e0130955. doi: 10.1371/journal.pone.0130955. eCollection 2015. PLoS One. 2015. PMID: 26161978 Free PMC article.
-
Uses of Staphylococcus aureus GeneChips in genotyping and genetic composition analysis.J Clin Microbiol. 2004 Sep;42(9):4275-83. doi: 10.1128/JCM.42.9.4275-4283.2004. J Clin Microbiol. 2004. PMID: 15365023 Free PMC article.
References
-
- Betley M J, Mekalanos J J. Staphylococcal enterotoxin A is encoded by phage. Science. 1985;229:185–187. - PubMed
-
- Bohach G A, Fast D J, Nelson R D, Schlievert P M. Staphylococcal and streptococcal pyrogenic toxins involved in toxic shock syndrome and related illnesses. Crit Rev Microbiol. 1990;17:251–272. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

