Molecular characterization of two natural hotspots in the Drosophila buzzatii genome induced by transposon insertions
- PMID: 11483576
- PMCID: PMC311088
- DOI: 10.1101/gr.174001
Molecular characterization of two natural hotspots in the Drosophila buzzatii genome induced by transposon insertions
Abstract
Transposable elements (TEs) have been implicated in the generation of genetic rearrangements, but their potential to mediate changes in the organization and architecture of host genomes could be even greater than previously thought. Here, we describe the naturally occurring structural and nucleotide variation around two TE insertions in the genome of Drosophila buzzatii. The studied regions correspond to the breakpoints of a widespread chromosomal inversion generated by ectopic recombination between oppositely oriented copies of a TE named Galileo. A detailed molecular analysis by Southern hybridization, PCR amplification, and DNA sequencing of 7.1 kb surrounding the inversion breakpoints in 39 D. buzzatii lines revealed an unprecedented degree of restructuring, consisting of 22 insertions of ten previously undescribed TEs, 13 deletions, 1 duplication, and 1 small inversion. All of these alterations occurred exclusively in inverted chromosomes and appear to have accumulated after the insertion of the Galileo elements, within or close to them. The nucleotide variation at the studied regions is six times lower in inverted than in noninverted chromosomes, suggesting that most of the observed changes originated in only 84,000 years. Galileo elements thus seemed to promote the transformation of these, otherwise normal, chromosomal regions in genetically unstable hotspots and highly efficient traps for transposon insertions. The particular features of two new Galileo copies found indicate that this TE belongs to the Foldback family. Together, our results strengthen the importance of TEs, and especially DNA transposons, as inducers of genome plasticity in evolution.
Figures



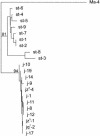
Comment in
-
Transposon-induced hotspots for genomic instability.Genome Res. 2001 Aug;11(8):1321-2. doi: 10.1101/gr.201201. Genome Res. 2001. PMID: 11483571 No abstract available.
References
-
- Adams MD, Celniker SE, Holt RA, Evans CA, Gocayne JD, Amanatides PG, Scherer SE, Li PW, Hoskins RA, Galle RF, et al. The genome sequence of Drosophila melanogaster. Science. 2000;287:2185–2195. - PubMed
-
- Albertini AM, Hofer M, Calos MP, Miller JH. On the formation of spontaneous deletions: The importance of short sequence homologies in the generation of large deletions. Cell. 1982;29:319–328. - PubMed
-
- Aquadro CF. Molecular population genetics of Drosophila. In: Oakeshott J, Whitten MJ, editors. Molecular approaches to fundamental and applied entomology. New York: Springer-Verlag; 1993. pp. 222–266.
-
- Arkhipova IR, Lyubomirskaya NV, Ilyin YV. Drosophila retrotransposons. Heidelberg: Springer-Verlag; 1995.
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
